TF_ChIP-seq_ENCSR487LUQ
| Id: | TF_ChIP-seq/ENCSR487LUQ |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR487LUQ [biosamplesummary="Homo sapiens HepG2" and target="PHF8"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN307CFQ|/analyses/ENCAN307CFQ/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF327PCB|/files/ENCFF327PCB/}, {ENCFF060MLS|/files/ENCFF060MLS/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 1.29 and a self consistency ratio of 4.47. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF796PPS|/files/ENCFF796PPS/}, {ENCFF661RKT|/files/ENCFF661RKT/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 1.29 and a self consistency ratio of 4.47. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
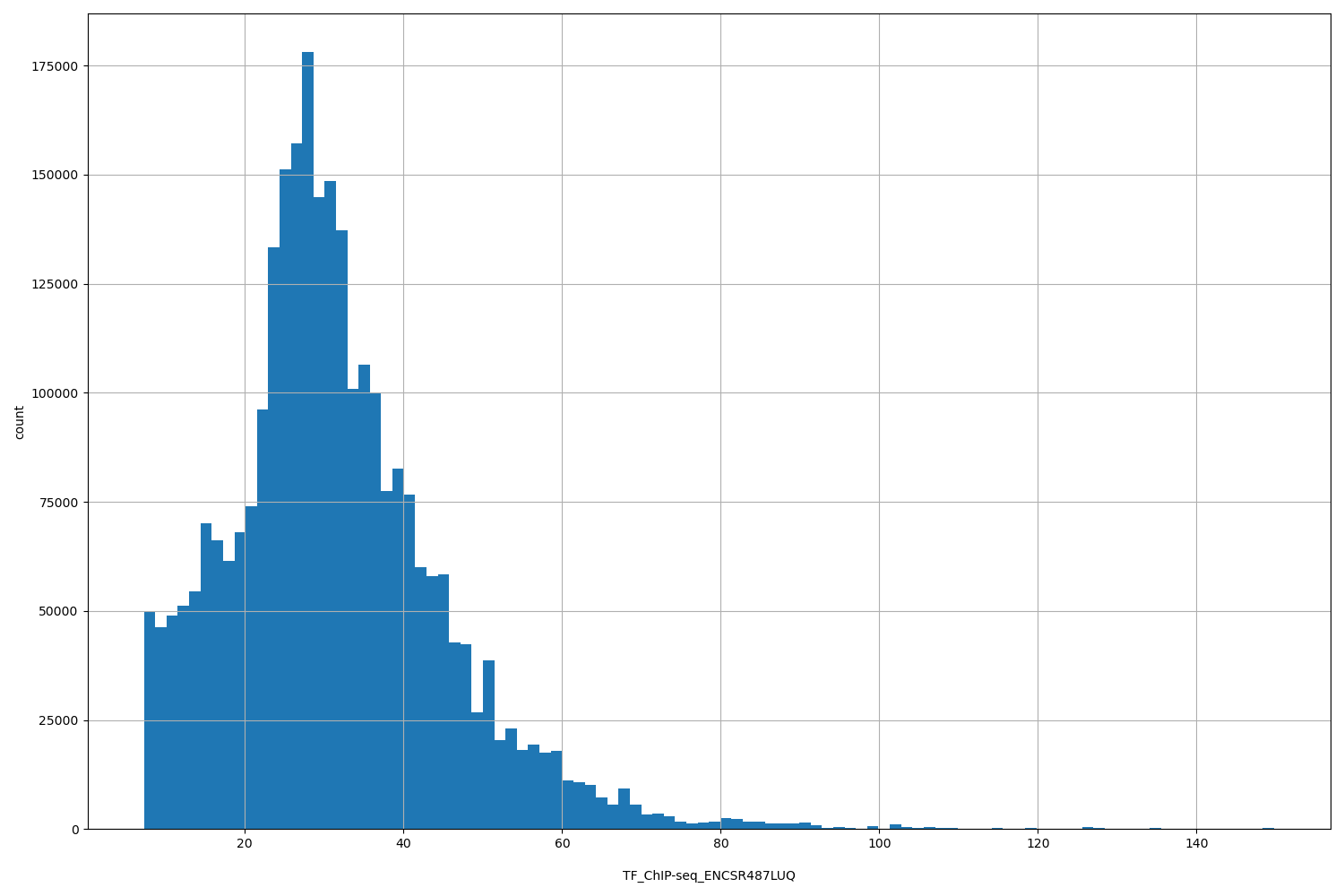
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR487LUQ | float |
TF_ChIP-seq_ENCSR487LUQ |
TF_ChIP-seq ENCSR487LUQ [biosample_summary="Homo sapiens HepG2" and target="PHF8"]
|
 |
[7.26, 150] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF060MLS.bed.gz | 106.62 KB | 52abfc24697bfc447f22344fa5dba7dd |
| ENCFF060MLS.bed.gz.dvc | 100.0 B | 8668f29fbff9aed4b9ad75108bdb9233 |
| ENCFF060MLS.tabix.bed.gz | 84.73 KB | 62287bcbd4045dd74ba9fe62752e2d7e |
| ENCFF060MLS.tabix.bed.gz.dvc | 105.0 B | 9dd963542e8f574a07d446b60ff8da9e |
| ENCFF060MLS.tabix.bed.gz.tbi | 45.82 KB | cdef902351785589c7cff3afdf129ca2 |
| ENCFF060MLS.tabix.bed.gz.tbi.dvc | 109.0 B | d5f0911a13d3ab4fdc7425c447283327 |
| genomic_resource.yaml | 2.47 KB | dee3b7a60ecdb53eb9ad050f1eaeedff |
| genomic_resource_original.yaml | 2.38 KB | e536675362ef1671ea3ec5b7e136c283 |
| statistics/ |