TF_ChIP-seq_ENCSR449SEF
| Id: | TF_ChIP-seq/ENCSR449SEF |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR449SEF [biosamplesummary="Homo sapiens transverse colon tissue female adult (51 years)" and target="CTCF"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: female adult (51 years) output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN830XVY|/analyses/ENCAN830XVY/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF119LFI|/files/ENCFF119LFI/}, {ENCFF986BNJ|/files/ENCFF986BNJ/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 1.20 and a self consistency ratio of 2.06. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF406OWW|/files/ENCFF406OWW/}, {ENCFF961BPY|/files/ENCFF961BPY/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 1.20 and a self consistency ratio of 2.06. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
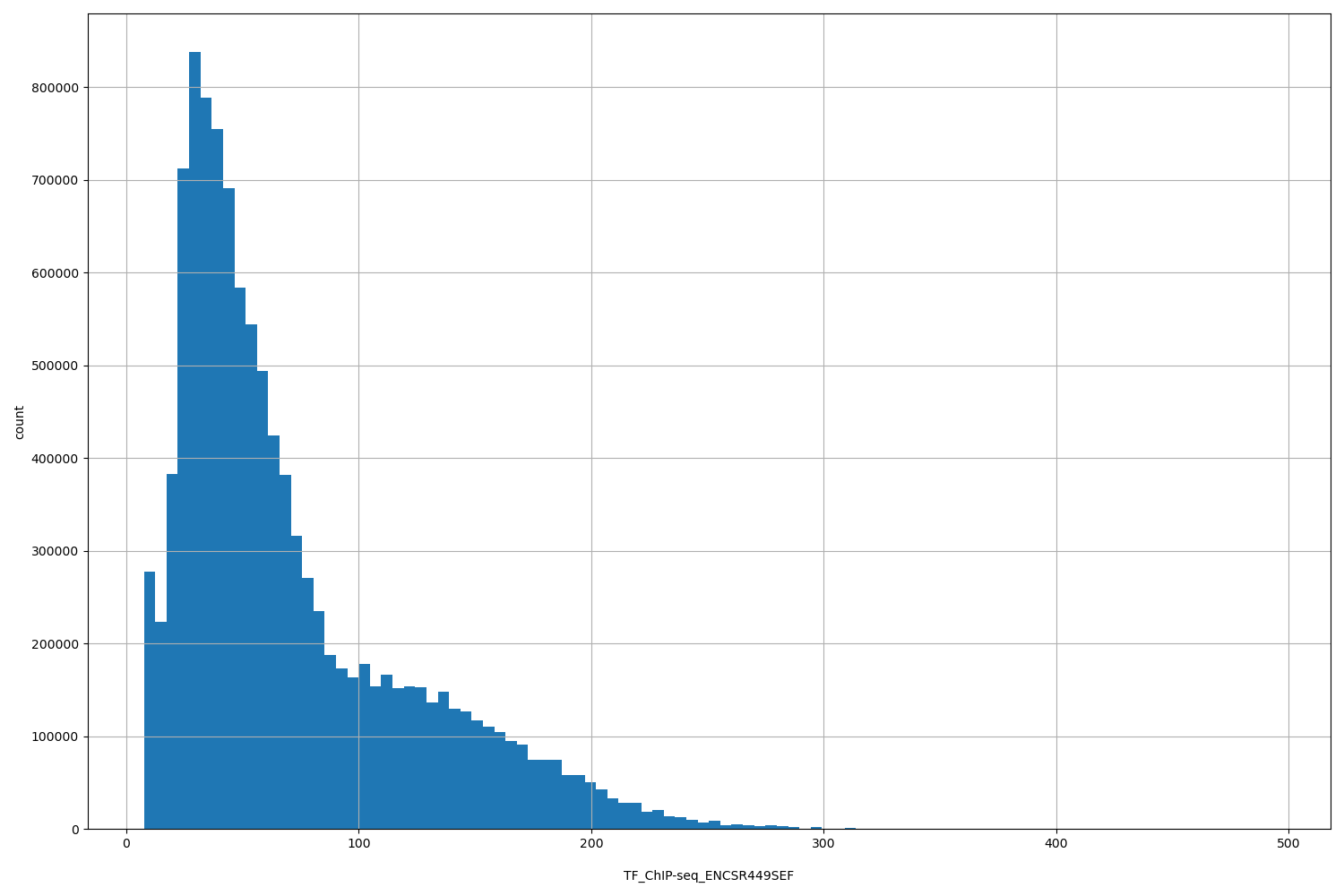
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR449SEF | float |
TF_ChIP-seq_ENCSR449SEF |
TF_ChIP-seq ENCSR449SEF [biosample_summary="Homo sapiens transverse colon tissue female adult (51 years)" and target="CTCF"]
|
 |
[7.54, 494] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF406OWW.bed.gz | 523.0 KB | b5c3089cebf64bb398905a09268e07dd |
| ENCFF406OWW.bed.gz.dvc | 100.0 B | 6b7842b03ad502c904d6f91facc10e25 |
| ENCFF406OWW.tabix.bed.gz | 406.4 KB | 7155b010df97fe715fe853112cbd0a7c |
| ENCFF406OWW.tabix.bed.gz.dvc | 106.0 B | eac3080d864a73a98f9617cb0872b747 |
| ENCFF406OWW.tabix.bed.gz.tbi | 206.16 KB | 65bf9cde3d609b71190cc23bbe56b347 |
| ENCFF406OWW.tabix.bed.gz.tbi.dvc | 110.0 B | b021124e0626b8c31be855bbb423f65b |
| genomic_resource.yaml | 2.59 KB | 9d8a64d860b5f391a03aa60bcbb52844 |
| genomic_resource_original.yaml | 2.46 KB | 9598dcc26608cbbf42d33e5e08359801 |
| statistics/ |