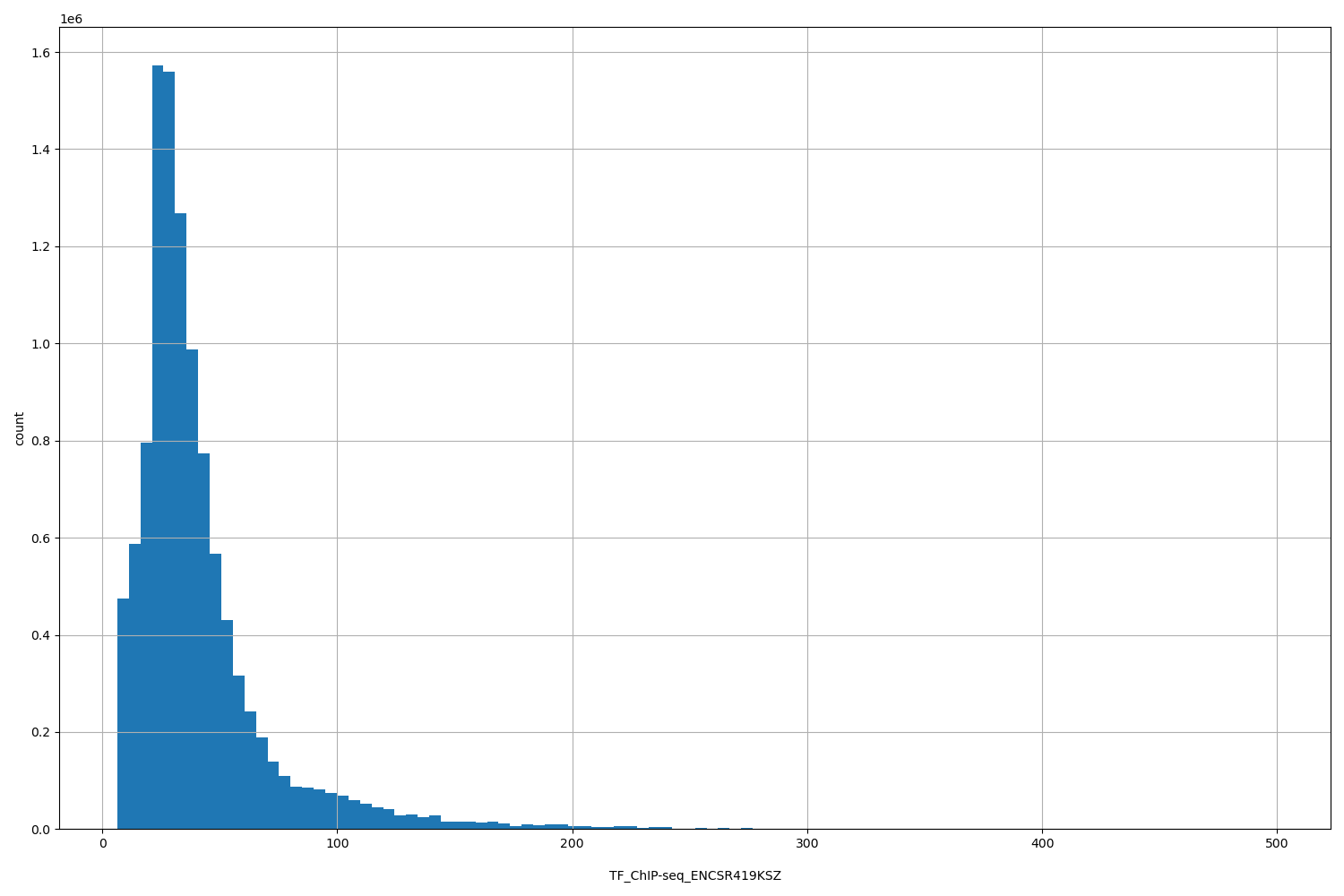
TF_ChIP-seq_ENCSR419KSZ
| Id: | TF_ChIP-seq/ENCSR419KSZ |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR419KSZ [biosamplesummary="Homo sapiens HepG2" and target="NCOR1"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN755IMV|/analyses/ENCAN755IMV/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Processed alignments file {ENCFF707EHW|/files/ENCFF707EHW/} processed by ChIP-seq ENCODE3 hg19 pipeline has 19963828 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting NCOR1-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF193YSE|/files/ENCFF193YSE/}, {ENCFF033BDT|/files/ENCFF033BDT/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 1.64 and a self consistency ratio of 4.62. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF874TMI|/files/ENCFF874TMI/}, {ENCFF754GAA|/files/ENCFF754GAA/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 1.64 and a self consistency ratio of 4.62. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR419KSZ | float |
TF_ChIP-seq_ENCSR419KSZ |
TF_ChIP-seq ENCSR419KSZ [biosample_summary="Homo sapiens HepG2" and target="NCOR1"]
|
 |
[6.34, 498] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF033BDT.bed.gz | 416.0 KB | adb45f94576c907fd380d8a5a4defd4a |
| ENCFF033BDT.bed.gz.dvc | 100.0 B | 95d6d7f2824bbc16c30c6cb4741895a1 |
| ENCFF033BDT.tabix.bed.gz | 337.71 KB | 904d9d5e21ec6d5bcf749fb428abef89 |
| ENCFF033BDT.tabix.bed.gz.dvc | 106.0 B | 36cdfa650a7d4e8b1d0ebb694ab031ca |
| ENCFF033BDT.tabix.bed.gz.tbi | 130.82 KB | 44d03114ac6aeb715fa9316554558bcf |
| ENCFF033BDT.tabix.bed.gz.tbi.dvc | 110.0 B | 7e64efd043df2f049debae6e52c884ef |
| genomic_resource.yaml | 2.97 KB | 864ca6cc1418510f355fc3a5a6441222 |
| genomic_resource_original.yaml | 2.87 KB | b39d93001d4e34e6658e0dadb9daf5ed |
| statistics/ |