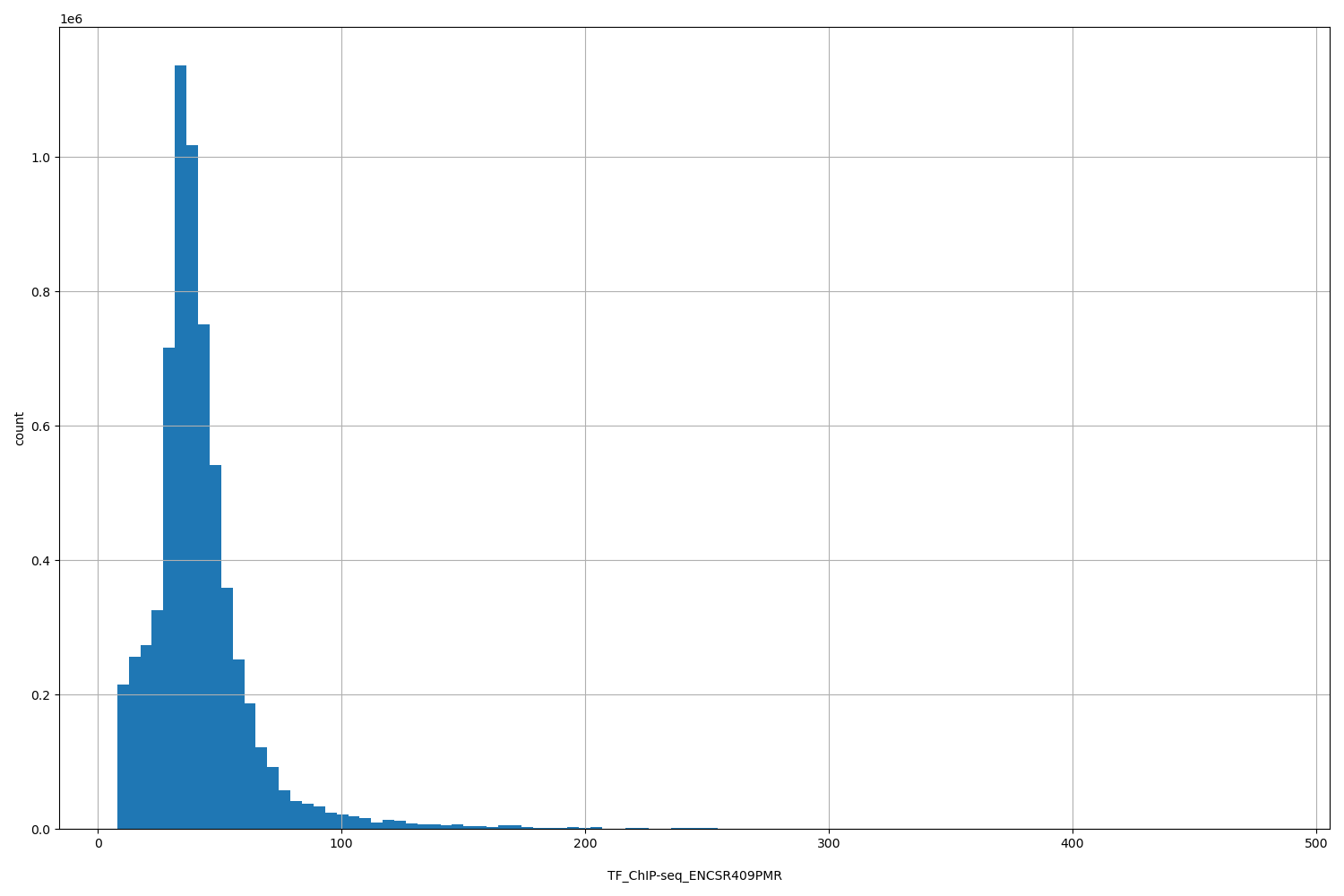
TF_ChIP-seq_ENCSR409PMR
| Id: | TF_ChIP-seq/ENCSR409PMR |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR409PMR [biosamplesummary="Homo sapiens HepG2 genetically modified (insertion) using CRISPR targeting H. sapiens ZBED4" and target="ZBED4"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: genetically modified (insertion) using CRISPR targeting H. sapiens ZBED4 output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN412LCF|/analyses/ENCAN412LCF/} has in progress subobject document {a44fb172-cbf1-4538-b25e-9b222c94ec17|/documents/a44fb172-cbf1-4538-b25e-9b222c94ec17/} audit_internal_action: Released analysis {ENCAN412LCF|/analyses/ENCAN412LCF/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF021YBN|/files/ENCFF021YBN/} processed by ChIP-seq ENCODE4 v1.4.0 GRCh38 pipeline have a rescue ratio of 1.10 and a self consistency ratio of 2.46. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF157CDZ|/files/ENCFF157CDZ/} processed by ChIP-seq ENCODE4 v1.4.0 GRCh38 pipeline have a rescue ratio of 1.10 and a self consistency ratio of 2.46. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR409PMR | float |
TF_ChIP-seq_ENCSR409PMR |
TF_ChIP-seq ENCSR409PMR [biosample_summary="Homo sapiens HepG2 genetically modified (insertion) using CRISPR targeting H. sapiens ZBED4" and target="ZBED4"]
|
 |
[7.97, 482] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF021YBN.bed.gz | 283.96 KB | ec5b13756a26bfb4081c7dc1a2d31e33 |
| ENCFF021YBN.bed.gz.dvc | 100.0 B | 9372e83badc64baddb61b1ac529203ad |
| ENCFF021YBN.tabix.bed.gz | 218.71 KB | 75082f64e665bb47ff5cbb6a9e92f6c4 |
| ENCFF021YBN.tabix.bed.gz.dvc | 106.0 B | 03ecc287f8401a6bc41ecf4ec631600e |
| ENCFF021YBN.tabix.bed.gz.tbi | 103.31 KB | 1de50265c3ff59116312af69067ec051 |
| ENCFF021YBN.tabix.bed.gz.tbi.dvc | 110.0 B | e1d230e3868d081bbd97443560756bd8 |
| genomic_resource.yaml | 2.95 KB | d24ed9a8400b98012784b4221142d7f4 |
| genomic_resource_original.yaml | 2.79 KB | cc0fba7139eec7d4bbf7f3b685a07343 |
| statistics/ |