TF_ChIP-seq_ENCSR407BEZ
| Id: | TF_ChIP-seq/ENCSR407BEZ |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR407BEZ [biosamplesummary="Homo sapiens HepG2" and target="ZHX2"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN367ORJ|/analyses/ENCAN367ORJ/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Processed alignments file {ENCFF162VTZ|/files/ENCFF162VTZ/} processed by ChIP-seq ENCODE3 hg19 pipeline has 17952236 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting ZHX2-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
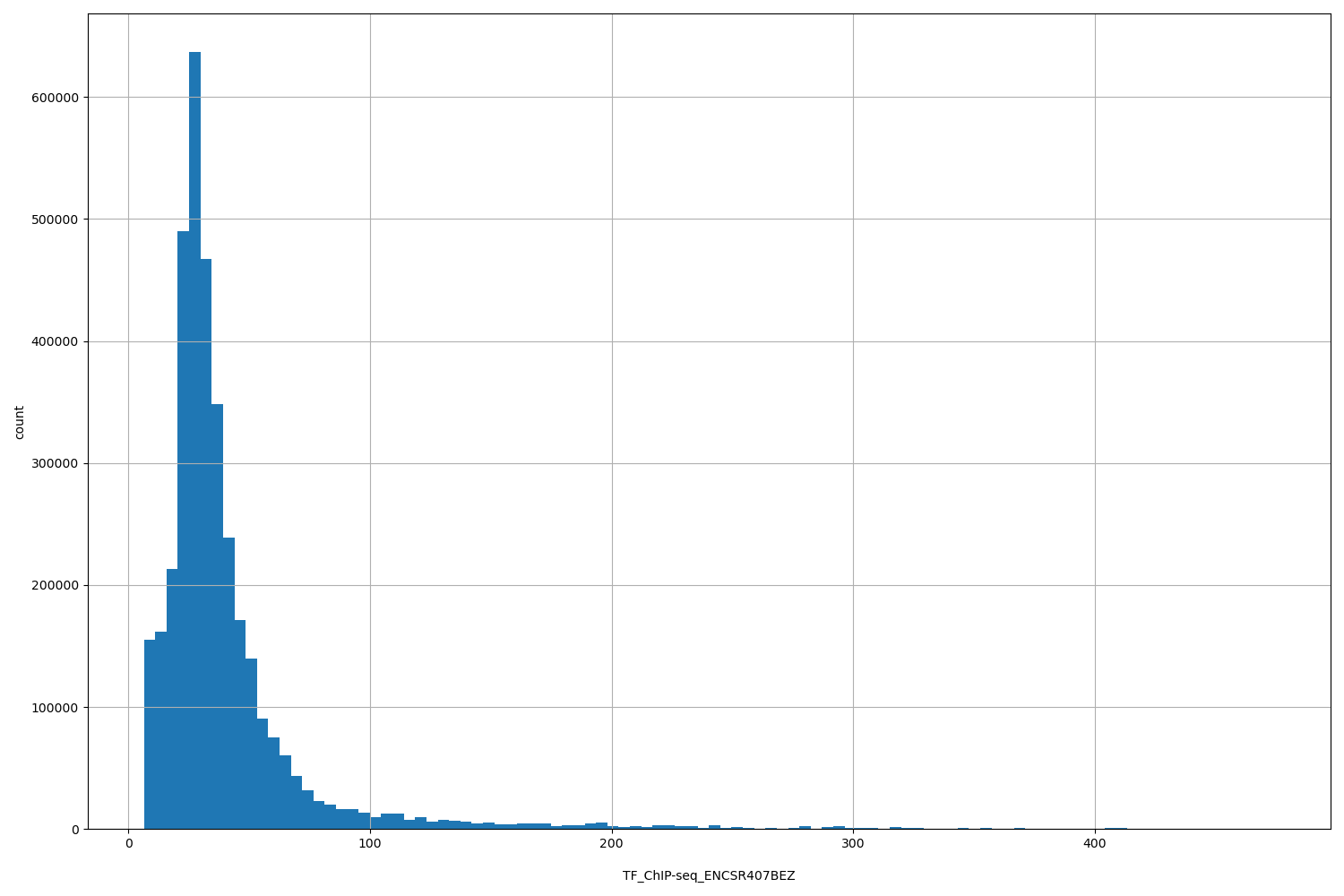
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR407BEZ | float |
TF_ChIP-seq_ENCSR407BEZ |
TF_ChIP-seq ENCSR407BEZ [biosample_summary="Homo sapiens HepG2" and target="ZHX2"]
|
 |
[6.49, 474] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF974OIC.bed.gz | 177.06 KB | 79e70a2005e90b9615aa7013c7effab6 |
| ENCFF974OIC.bed.gz.dvc | 100.0 B | 53d5b58a3d0db961a6db27ac3c2e2fd8 |
| ENCFF974OIC.tabix.bed.gz | 136.61 KB | ecb566adee967014f1309810e842c512 |
| ENCFF974OIC.tabix.bed.gz.dvc | 106.0 B | 95b710df15db7d1ed149c2d3cd0e95a6 |
| ENCFF974OIC.tabix.bed.gz.tbi | 77.31 KB | 538929e6fc1b6d5d6ebafa337459dd5d |
| ENCFF974OIC.tabix.bed.gz.tbi.dvc | 109.0 B | 25d90bbace6a67933536f82b6d1520a5 |
| genomic_resource.yaml | 1.83 KB | f6f0f8f1fa25001925728177aa8a22f9 |
| genomic_resource_original.yaml | 1.74 KB | 5cbfee744bee13407a622eaf60e2594a |
| statistics/ |