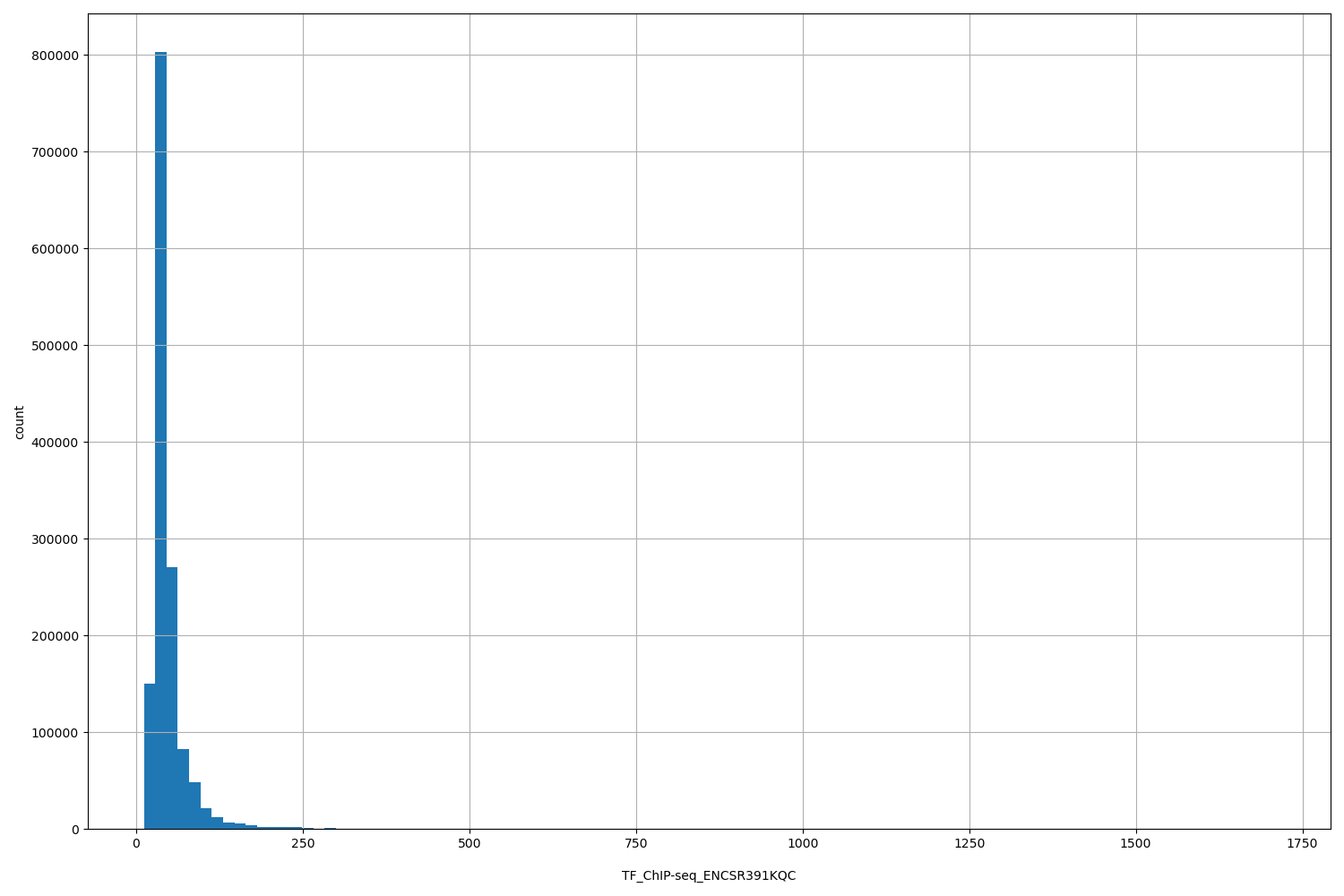
TF_ChIP-seq_ENCSR391KQC
| Id: | TF_ChIP-seq/ENCSR391KQC |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR391KQC [biosamplesummary="Homo sapiens MCF-7" and target="MTA3"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN981OEJ|/analyses/ENCAN981OEJ/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_internal_action: File {ENCFF083AZM|/files/ENCFF083AZM/} with status 'released' is derived from file {ENCFF419RSJ|/files/ENCFF419RSJ/} with status 'archived'. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF083AZM|/files/ENCFF083AZM/}, {ENCFF384YHL|/files/ENCFF384YHL/} processed by ChIP-seq ENCODE3 GRCh38 pipeline have a rescue ratio of 2.42 and a self consistency ratio of 1.40. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF668JOA|/files/ENCFF668JOA/}, {ENCFF549VFF|/files/ENCFF549VFF/} processed by ChIP-seq ENCODE3 GRCh38 pipeline have a rescue ratio of 2.42 and a self consistency ratio of 1.40. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR391KQC | float |
TF_ChIP-seq_ENCSR391KQC |
TF_ChIP-seq ENCSR391KQC [biosample_summary="Homo sapiens MCF-7" and target="MTA3"]
|
 |
[11.6, 1.71e+03] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF083AZM.bed.gz | 81.13 KB | 309de1dcaf93091600635d9a08bdfaf2 |
| ENCFF083AZM.bed.gz.dvc | 99.0 B | 709d6a8724b42bb5c73570c2c9208680 |
| ENCFF083AZM.tabix.bed.gz | 54.56 KB | b32195eb18b7cc31f82c95ea19b3a041 |
| ENCFF083AZM.tabix.bed.gz.dvc | 105.0 B | 2fbddd5d2c82cb9e2759cb17bbf98aa0 |
| ENCFF083AZM.tabix.bed.gz.tbi | 37.49 KB | 6de9dee90e4cd7f6e3f29719831847d7 |
| ENCFF083AZM.tabix.bed.gz.tbi.dvc | 109.0 B | 7190605615b9faa0f7a083069e882431 |
| genomic_resource.yaml | 2.66 KB | 8f647d05c166716785ba16a20bff6634 |
| genomic_resource_original.yaml | 2.57 KB | b2c99e052d2fe0e20bb0421b5f51281c |
| statistics/ |