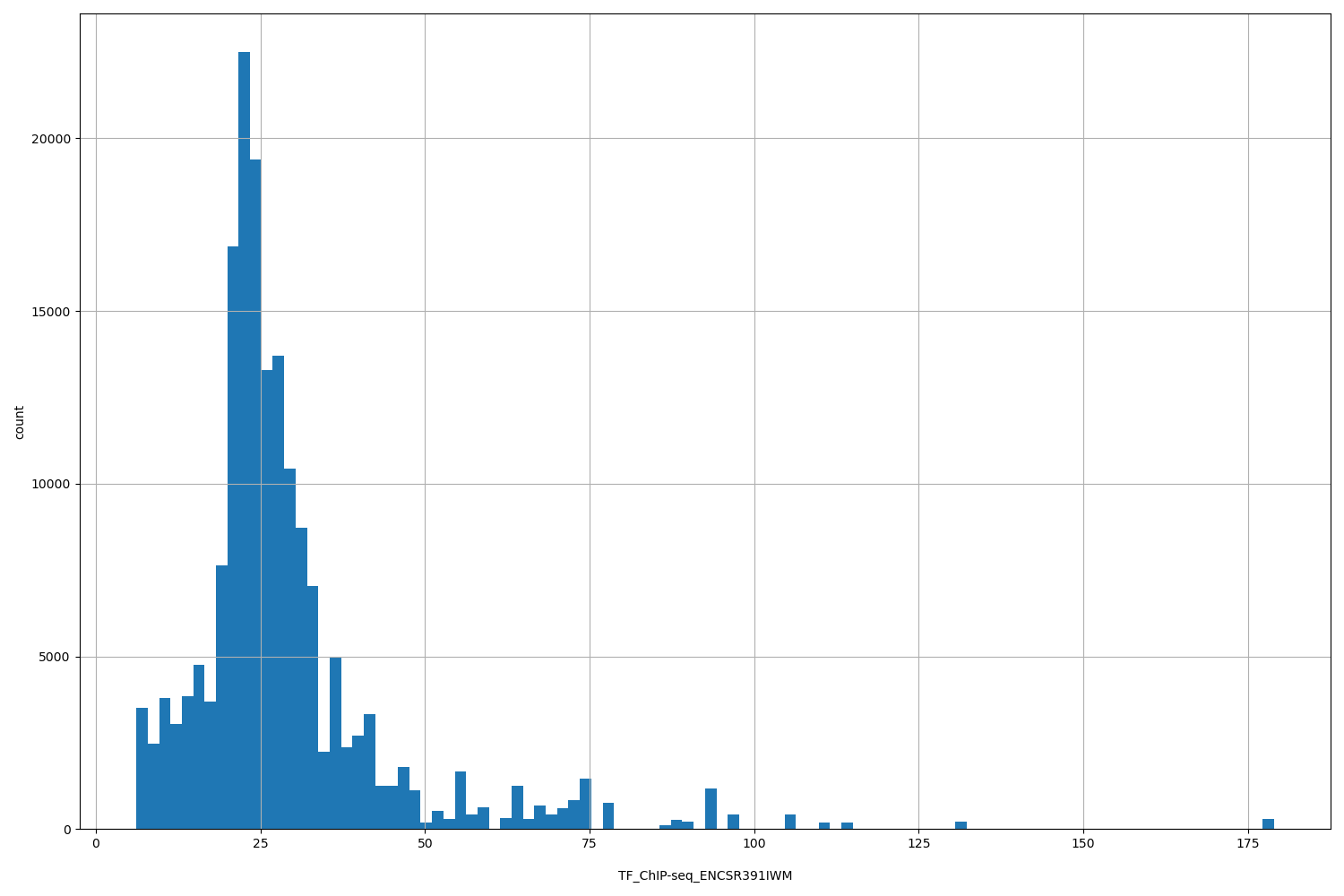
TF_ChIP-seq_ENCSR391IWM
| Id: | TF_ChIP-seq/ENCSR391IWM |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR391IWM [biosamplesummary="Homo sapiens GM12878" and target="KDM1A"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN289BRH|/analyses/ENCAN289BRH/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_internal_action: File {ENCFF799KZP|/files/ENCFF799KZP/} with status 'released' is derived from file {ENCFF419RSJ|/files/ENCFF419RSJ/} with status 'archived'. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF799KZP|/files/ENCFF799KZP/}, {ENCFF355CKA|/files/ENCFF355CKA/} processed by ChIP-seq ENCODE3 GRCh38 pipeline have a rescue ratio of 1.30 and a self consistency ratio of 5.34. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF252WSP|/files/ENCFF252WSP/}, {ENCFF423NOY|/files/ENCFF423NOY/} processed by ChIP-seq ENCODE3 GRCh38 pipeline have a rescue ratio of 1.30 and a self consistency ratio of 5.34. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR391IWM | float |
TF_ChIP-seq_ENCSR391IWM |
TF_ChIP-seq ENCSR391IWM [biosample_summary="Homo sapiens GM12878" and target="KDM1A"]
|
 |
[6.14, 179] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF799KZP.bed.gz | 8.72 KB | 44ff40f3cbbb5942096b3bacc95ae072 |
| ENCFF799KZP.bed.gz.dvc | 98.0 B | 9826a81ed2edf76fd5cf06aadc800954 |
| ENCFF799KZP.tabix.bed.gz | 6.16 KB | d62c035224ad1fcfa6954ddb019a3b5f |
| ENCFF799KZP.tabix.bed.gz.dvc | 104.0 B | ee1ae9e0aa86d45e7189ba8baac7eaed |
| ENCFF799KZP.tabix.bed.gz.tbi | 8.49 KB | 9f53b1633f62141843aefd8a8a11badf |
| ENCFF799KZP.tabix.bed.gz.tbi.dvc | 108.0 B | 31fa1b49a113f53326019aa9df49104d |
| genomic_resource.yaml | 2.67 KB | ad53c0366074716d6d929ed96f5418b7 |
| genomic_resource_original.yaml | 2.57 KB | 1e992991b11b82c5cab6b8bccaa2231c |
| statistics/ |