TF_ChIP-seq_ENCSR370NFS
| Id: | TF_ChIP-seq/ENCSR370NFS |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR370NFS [biosamplesummary="Homo sapiens K562" and target="ZNF280A"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN512XGK|/analyses/ENCAN512XGK/} has in progress subobject document {a8593378-5e8c-458a-9afe-6d783dc7e156|/documents/a8593378-5e8c-458a-9afe-6d783dc7e156/} audit_internal_action: Released analysis {ENCAN512XGK|/analyses/ENCAN512XGK/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_warning: Processed alignments file {ENCFF105QHT|/files/ENCFF105QHT/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 14262369 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting ZNF280A-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Processed alignments file {ENCFF061PEU|/files/ENCFF061PEU/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 14668609 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting ZNF280A-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: PBC1 (PCR Bottlenecking Coefficient 1, M1/M_distinct) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where some reads map (M_distinct). A PBC1 value in the range 0 - 0.5 is severe bottlenecking, 0.5 - 0.8 is moderate bottlenecking, 0.8 - 0.9 is mild bottlenecking, and > 0.9 is no bottlenecking. PBC1 value > 0.9 is recommended, but > 0.8 is acceptable. ENCODE processed alignments file {ENCFF105QHT|/files/ENCFF105QHT/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline was generated from a library with PBC1 value of 0.90. audit_warning: PBC2 (PCR Bottlenecking Coefficient 2, M1/M2) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where two reads map uniquely (M2). A PBC2 value in the range 0 - 1 is severe bottlenecking, 1 - 3 is moderate bottlenecking, 3 - 10 is mild bottlenecking, > 10 is no bottlenecking. PBC2 value > 10 is recommended, but > 3 is acceptable. ENCODE processed alignments file {ENCFF105QHT|/files/ENCFF105QHT/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline was generated from a library with PBC2 value of 9.81. |
| Labels: |
|
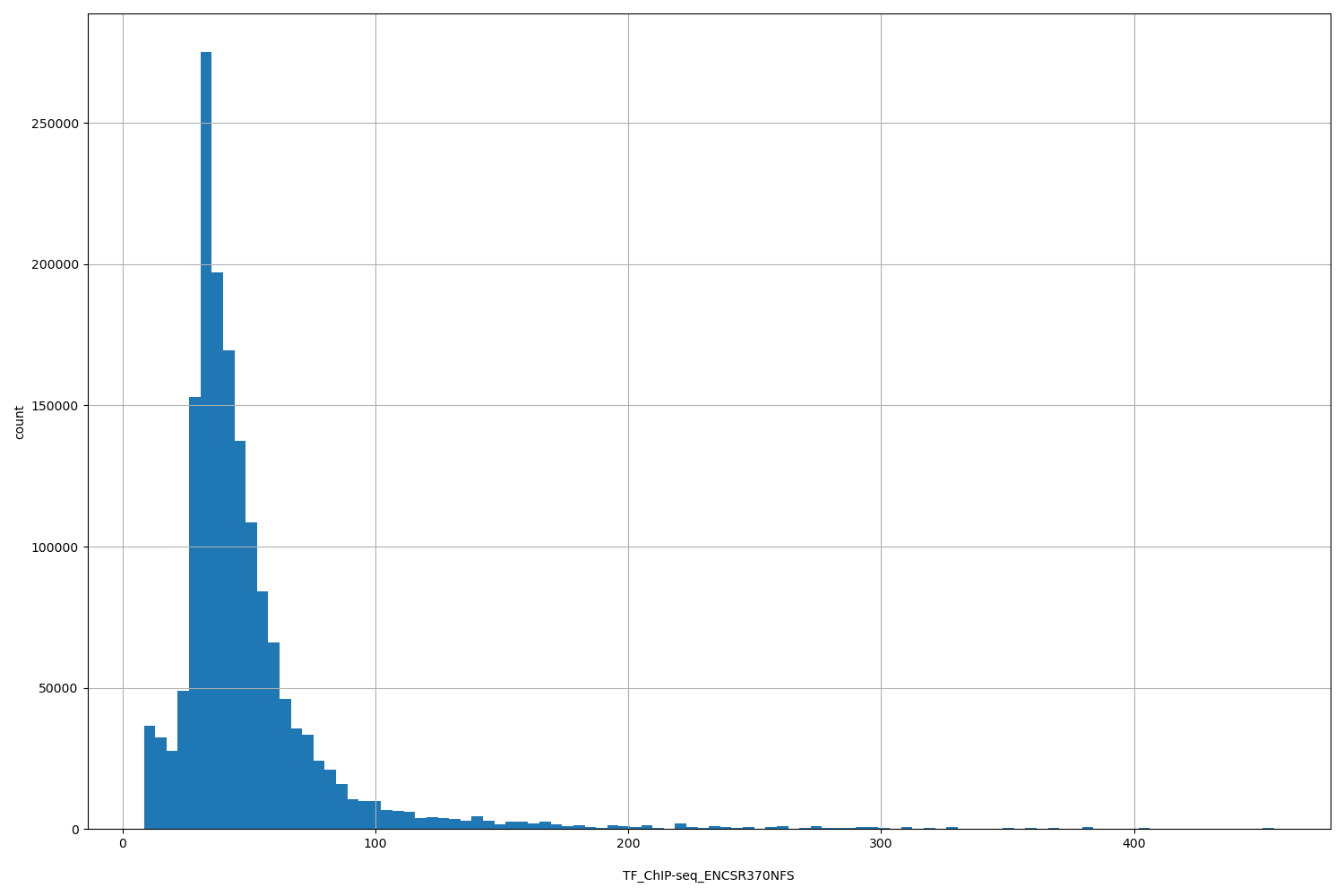
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR370NFS | float |
TF_ChIP-seq_ENCSR370NFS |
TF_ChIP-seq ENCSR370NFS [biosample_summary="Homo sapiens K562" and target="ZNF280A"]
|
 |
[8.52, 455] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF932ZCO.bed.gz | 92.54 KB | 4b9dd2b0782220dd498c0275d005f4ad |
| ENCFF932ZCO.bed.gz.dvc | 99.0 B | 469af4de34616ab918623d11fff27ccd |
| ENCFF932ZCO.tabix.bed.gz | 62.69 KB | 1880e8bfbc3b6c1ce5bc0222d856e7df |
| ENCFF932ZCO.tabix.bed.gz.dvc | 105.0 B | 8818dd6701ce7079e7de6dde60323f73 |
| ENCFF932ZCO.tabix.bed.gz.tbi | 46.17 KB | bddd38ad191c232100d74e65b671ab1e |
| ENCFF932ZCO.tabix.bed.gz.tbi.dvc | 109.0 B | 74564d757034418b66ca9b419dc0507f |
| genomic_resource.yaml | 4.43 KB | 22e19ec6e7e05135cc4695dde9e84af2 |
| genomic_resource_original.yaml | 4.33 KB | 1deb74b3dee3e00619e86f7d3e317e7c |
| statistics/ |