TF_ChIP-seq_ENCSR367KYL
| Id: | TF_ChIP-seq/ENCSR367KYL |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR367KYL [biosamplesummary="Homo sapiens HepG2 genetically modified (insertion) using CRISPR targeting H. sapiens ZHX3" and target="ZHX3"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: genetically modified (insertion) using CRISPR targeting H. sapiens ZHX3 output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN911FCT|/analyses/ENCAN911FCT/} has in progress subobject document {b924a4ae-59f4-4cd0-b265-c6e95080244f|/documents/b924a4ae-59f4-4cd0-b265-c6e95080244f/} audit_internal_action: Released analysis {ENCAN911FCT|/analyses/ENCAN911FCT/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF438MZI|/files/ENCFF438MZI/} processed by ChIP-seq ENCODE4 v1.4.0 GRCh38 pipeline have a rescue ratio of 1.45 and a self consistency ratio of 5.17. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF631YWI|/files/ENCFF631YWI/} processed by ChIP-seq ENCODE4 v1.4.0 GRCh38 pipeline have a rescue ratio of 1.45 and a self consistency ratio of 5.17. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
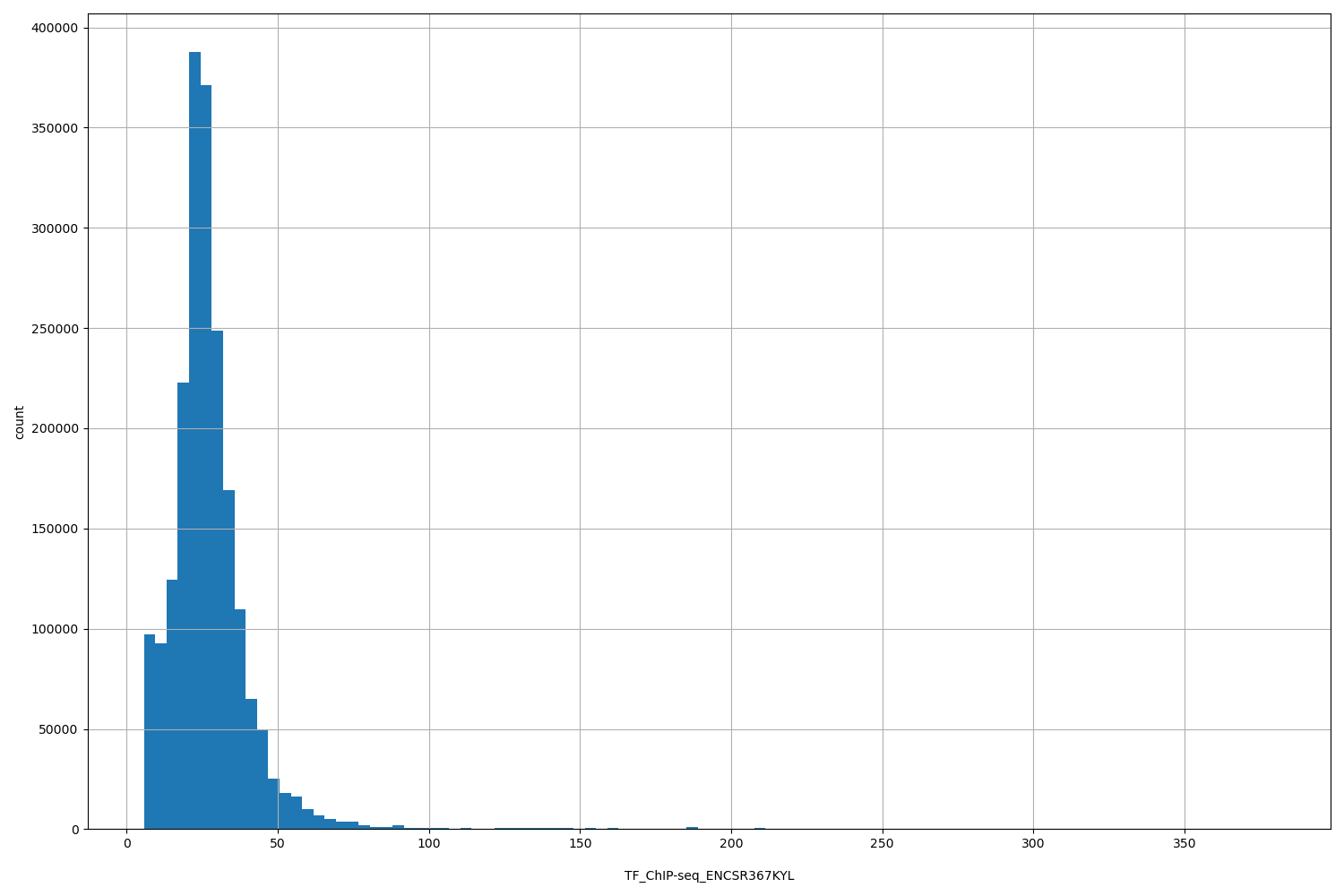
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR367KYL | float |
TF_ChIP-seq_ENCSR367KYL |
TF_ChIP-seq ENCSR367KYL [biosample_summary="Homo sapiens HepG2 genetically modified (insertion) using CRISPR targeting H. sapiens ZHX3" and target="ZHX3"]
|
 |
[5.74, 380] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF438MZI.bed.gz | 112.12 KB | 19b003bb2163d4d4de34a0f6913d15f7 |
| ENCFF438MZI.bed.gz.dvc | 100.0 B | ec9495fabd580e9e9fbf70d35e336c40 |
| ENCFF438MZI.tabix.bed.gz | 86.61 KB | 13e04e2eb130d55b794f78e35b104fd4 |
| ENCFF438MZI.tabix.bed.gz.dvc | 105.0 B | 79694ee2a1471f6eb53499a6449675d6 |
| ENCFF438MZI.tabix.bed.gz.tbi | 55.5 KB | 0d6eddbe1aafe99271478f1ea3d477e0 |
| ENCFF438MZI.tabix.bed.gz.tbi.dvc | 109.0 B | 2eeca668ebdb100295e5fde665f4ec45 |
| genomic_resource.yaml | 2.94 KB | c6d6b43b72b84b5258444aa3835793b2 |
| genomic_resource_original.yaml | 2.78 KB | 36066edd43412b711bf032fc7425e662 |
| statistics/ |