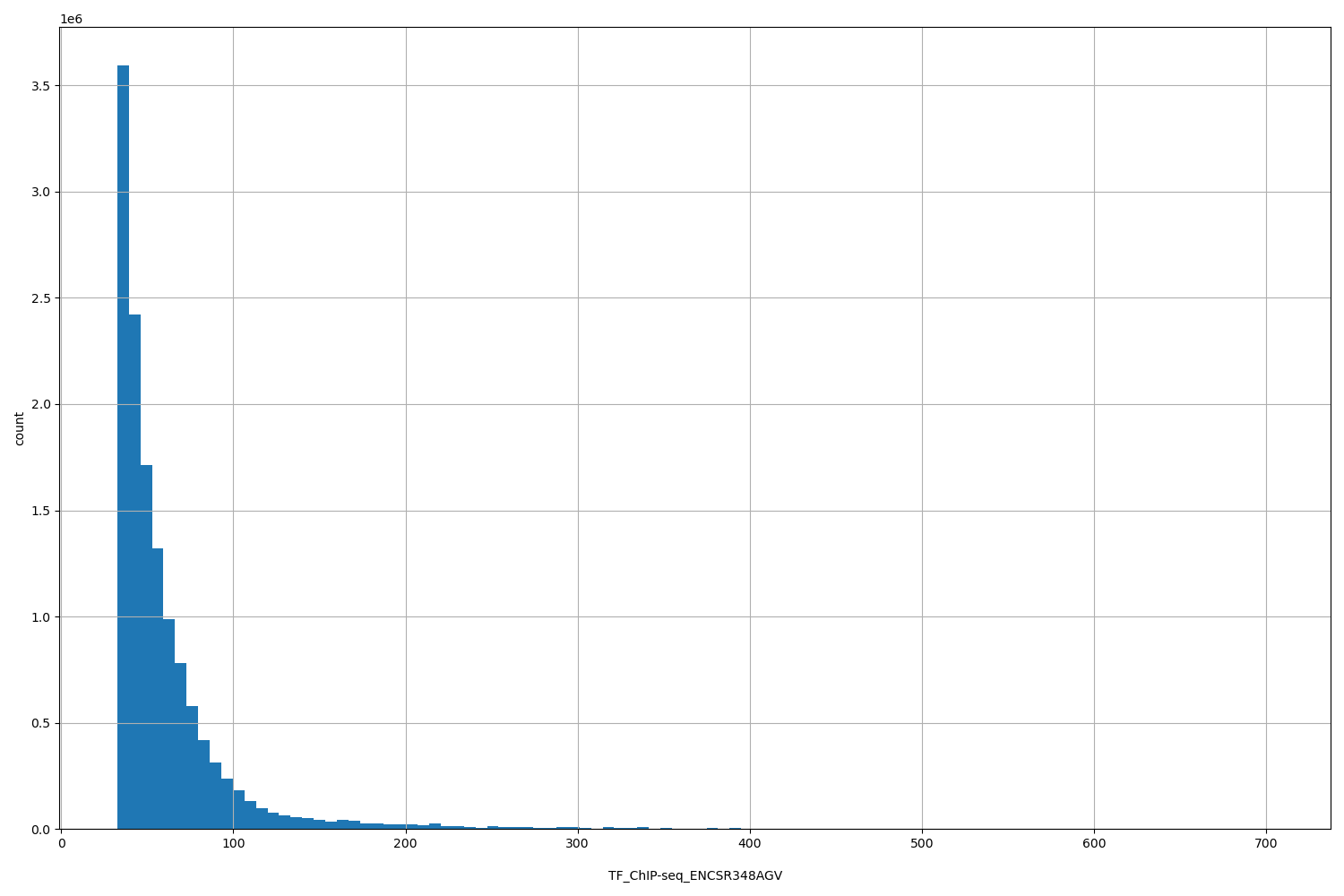
TF_ChIP-seq_ENCSR348AGV
TF_ChIP-seq ENCSR348AGV [biosample_summary="Homo sapiens HEK293" and target="SETDB1"]
| Id: | TF_ChIP-seq/ENCSR348AGV |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR348AGV [biosamplesummary="Homo sapiens HEK293" and target="SETDB1"] |
| Description: |
status: released biological_replicates: summary: output_type: optimal IDR thresholded peaks audit_internal_action: derived_from is a list of files that were used to create a given file; for example, fastq file(s) will appear in the derived_from list of an alignments file. Processed file {ENCFF002DAC|/files/ENCFF002DAC/} is missing the requisite file specification in its derived_from list. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR348AGV | float |
TF_ChIP-seq_ENCSR348AGV |
TF_ChIP-seq ENCSR348AGV [biosample_summary="Homo sapiens HEK293" and target="SETDB1"]
|
 |
[32.5, 704] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF002DAC.bed.gz | 449.9 KB | 5d5d24a11707ba48166091f5d4839bf0 |
| ENCFF002DAC.bed.gz.dvc | 100.0 B | a045244a9e499bb6d872774ff339d718 |
| ENCFF002DAC.tabix.bed.gz | 401.75 KB | a54258da5e56918da718a75863ff2d6f |
| ENCFF002DAC.tabix.bed.gz.dvc | 106.0 B | 044a951fd8b9c2902df5e7519268ab33 |
| ENCFF002DAC.tabix.bed.gz.tbi | 153.77 KB | 9e145e6f2a160f416427a81d507910e3 |
| ENCFF002DAC.tabix.bed.gz.tbi.dvc | 110.0 B | a13fcd7194dad8a1dc20116783fa1df2 |
| genomic_resource.yaml | 1.56 KB | 8b69ca724b840ff9c64346300db81942 |
| genomic_resource_original.yaml | 1.46 KB | 7ea32e3967f6a0a48d76b5801ca3dc00 |
| statistics/ |