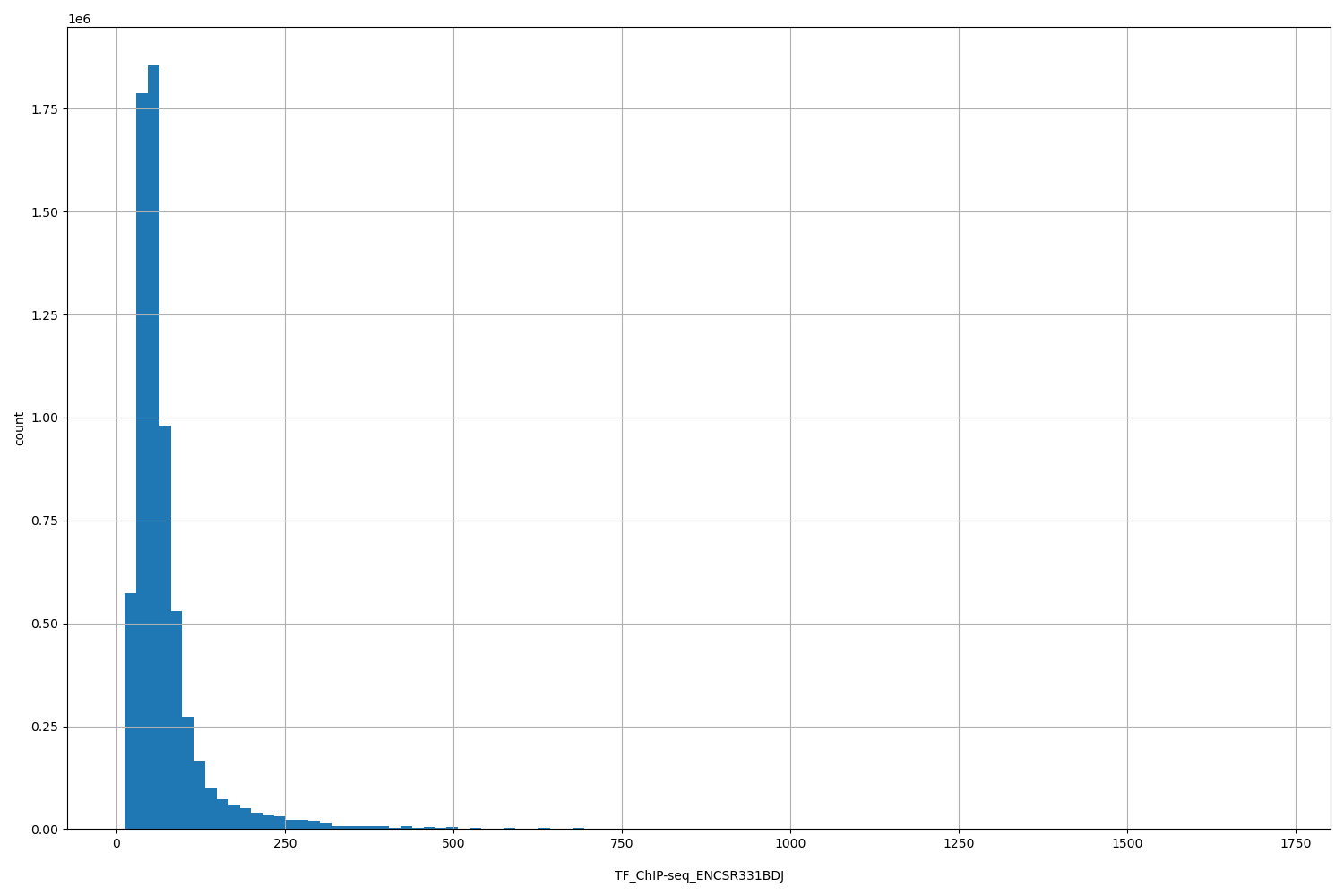
TF_ChIP-seq_ENCSR331BDJ
| Id: | TF_ChIP-seq/ENCSR331BDJ |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR331BDJ [biosamplesummary="Homo sapiens K562" and target="NKRF"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN712MYU|/analyses/ENCAN712MYU/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF596XXG|/files/ENCFF596XXG/}, {ENCFF856FSD|/files/ENCFF856FSD/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 1.38 and a self consistency ratio of 2.18. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF199DDB|/files/ENCFF199DDB/}, {ENCFF906PGD|/files/ENCFF906PGD/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 1.38 and a self consistency ratio of 2.18. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR331BDJ | float |
TF_ChIP-seq_ENCSR331BDJ |
TF_ChIP-seq ENCSR331BDJ [biosample_summary="Homo sapiens K562" and target="NKRF"]
|
 |
[12.7, 1.72e+03] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF906PGD.bed.gz | 325.74 KB | 1b3b69fc44e232cb151f753a73154afd |
| ENCFF906PGD.bed.gz.dvc | 100.0 B | 30f4517c42e074f0352245b24c3e205d |
| ENCFF906PGD.tabix.bed.gz | 237.0 KB | fef7d52ffa70bd660d667d1c2508a488 |
| ENCFF906PGD.tabix.bed.gz.dvc | 106.0 B | 087a2d92eec1af98d108fff80f9455e8 |
| ENCFF906PGD.tabix.bed.gz.tbi | 98.53 KB | 8a704f5b181bd5d7d39def326190943d |
| ENCFF906PGD.tabix.bed.gz.tbi.dvc | 110.0 B | 7cb33d4c337e22070bd098237ad82081 |
| genomic_resource.yaml | 2.47 KB | 703ac1d597b8d4daefc835d5b68bc99f |
| genomic_resource_original.yaml | 2.38 KB | d70c8419ed2efffa204abbcf1ea23906 |
| statistics/ |