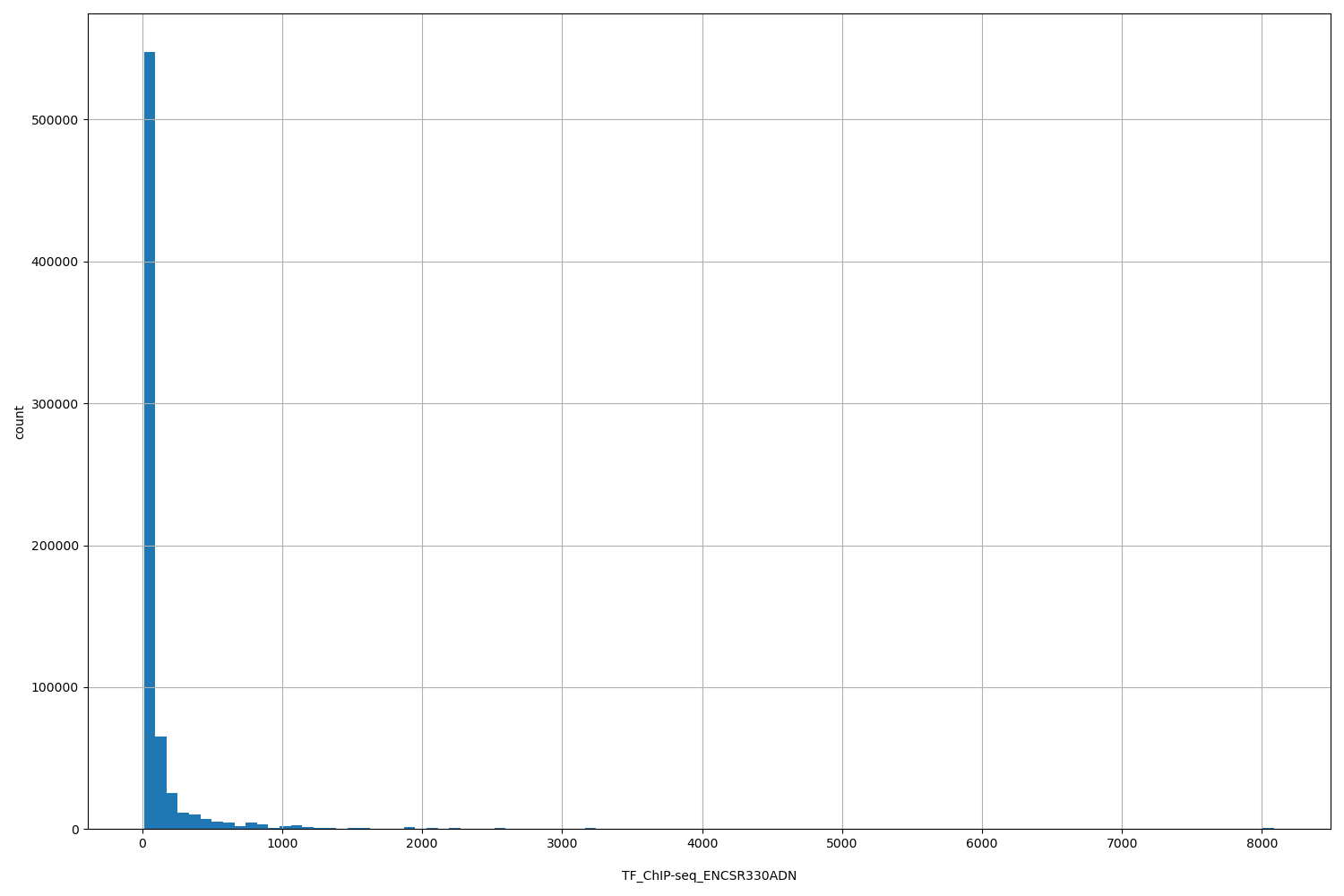
TF_ChIP-seq_ENCSR330ADN
| Id: | TF_ChIP-seq/ENCSR330ADN |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR330ADN [biosamplesummary="Homo sapiens MCF-7" and target="DDX20"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN170AXU|/analyses/ENCAN170AXU/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_internal_action: File {ENCFF089GNH|/files/ENCFF089GNH/} with status 'released' is derived from file {ENCFF419RSJ|/files/ENCFF419RSJ/} with status 'archived'. audit_warning: PBC1 (PCR Bottlenecking Coefficient 1, M1/M_distinct) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where some reads map (M_distinct). A PBC1 value in the range 0 - 0.5 is severe bottlenecking, 0.5 - 0.8 is moderate bottlenecking, 0.8 - 0.9 is mild bottlenecking, and > 0.9 is no bottlenecking. PBC1 value > 0.9 is recommended, but > 0.8 is acceptable. ENCODE processed alignments file {ENCFF682IXJ|/files/ENCFF682IXJ/} processed by ChIP-seq ENCODE3 GRCh38 pipeline was generated from a library with PBC1 value of 0.86. audit_warning: PBC2 (PCR Bottlenecking Coefficient 2, M1/M2) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where two reads map uniquely (M2). A PBC2 value in the range 0 - 1 is severe bottlenecking, 1 - 3 is moderate bottlenecking, 3 - 10 is mild bottlenecking, > 10 is no bottlenecking. PBC2 value > 10 is recommended, but > 3 is acceptable. ENCODE processed alignments file {ENCFF682IXJ|/files/ENCFF682IXJ/} processed by ChIP-seq ENCODE3 GRCh38 pipeline was generated from a library with PBC2 value of 6.83. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF089GNH|/files/ENCFF089GNH/}, {ENCFF558FCT|/files/ENCFF558FCT/} processed by ChIP-seq ENCODE3 GRCh38 pipeline have a rescue ratio of 1.62 and a self consistency ratio of 2.17. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF199QUM|/files/ENCFF199QUM/}, {ENCFF406DTF|/files/ENCFF406DTF/} processed by ChIP-seq ENCODE3 GRCh38 pipeline have a rescue ratio of 1.62 and a self consistency ratio of 2.17. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR330ADN | float |
TF_ChIP-seq_ENCSR330ADN |
TF_ChIP-seq ENCSR330ADN [biosample_summary="Homo sapiens MCF-7" and target="DDX20"]
|
 |
[12.1, 8.09e+03] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF089GNH.bed.gz | 56.85 KB | 9ad32a9822989a94e4bc03dc0f302059 |
| ENCFF089GNH.bed.gz.dvc | 99.0 B | def834670894ec3b40e4048fd41be8bc |
| ENCFF089GNH.tabix.bed.gz | 36.95 KB | 9f1455f9766f68796eb01b231f2a8a2b |
| ENCFF089GNH.tabix.bed.gz.dvc | 105.0 B | 58a363510cd0d053c319f5a64ea90077 |
| ENCFF089GNH.tabix.bed.gz.tbi | 29.99 KB | a3d9e4f7472cbaa0a19a2d53bcaf7073 |
| ENCFF089GNH.tabix.bed.gz.tbi.dvc | 109.0 B | c88251c347d4cf15cb907dc8313f701e |
| genomic_resource.yaml | 3.96 KB | 94c88bda73380fe97b29557f496d17ee |
| genomic_resource_original.yaml | 3.87 KB | dddbf92d3d5c3bcc6dfd99781b8b193e |
| statistics/ |