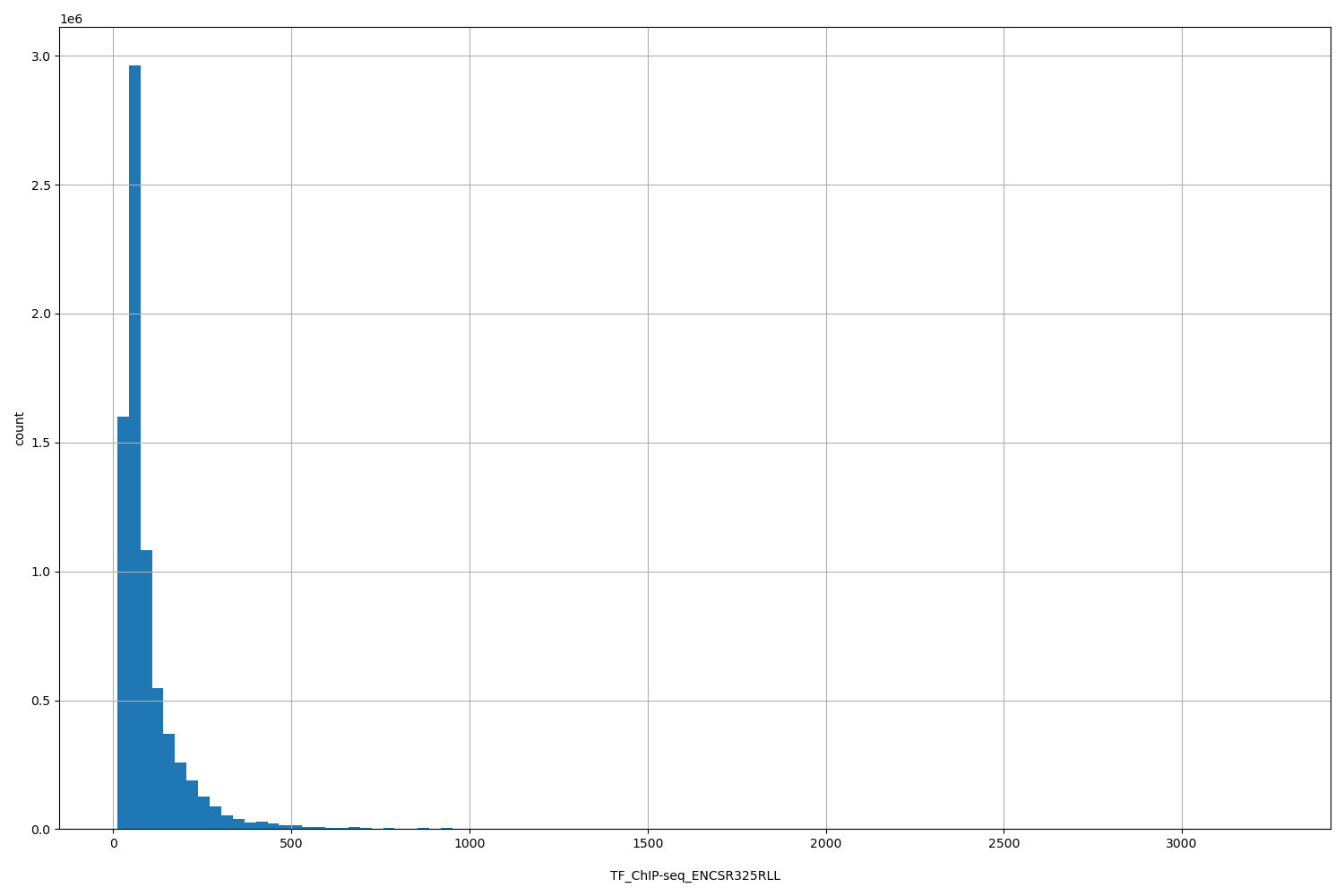
TF_ChIP-seq_ENCSR325RLL
| Id: | TF_ChIP-seq/ENCSR325RLL |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR325RLL [biosamplesummary="Homo sapiens K562" and target="POLR2B"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: optimal IDR thresholded peaks audit_internal_action: File {ENCFF078TXM|/files/ENCFF078TXM/} with status 'released' is derived from file {ENCFF419RSJ|/files/ENCFF419RSJ/} with status 'archived'. audit_internal_action: Archived analysis {ENCAN411CII|/analyses/ENCAN411CII/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Processed alignments file {ENCFF995XZH|/files/ENCFF995XZH/} processed by ChIP-seq ENCODE3 GRCh38 pipeline has 19503600 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting POLR2B-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR325RLL | float |
TF_ChIP-seq_ENCSR325RLL |
TF_ChIP-seq ENCSR325RLL [biosample_summary="Homo sapiens K562" and target="POLR2B"]
|
 |
[11.3, 3.26e+03] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF078TXM.bed.gz | 365.92 KB | cf30fcf78bc41a9d0703a890940edb6b |
| ENCFF078TXM.bed.gz.dvc | 100.0 B | cbda89c91fc01c35ced199257012354a |
| ENCFF078TXM.tabix.bed.gz | 260.4 KB | 3e9aa48e86912a828d9d04123707ecca |
| ENCFF078TXM.tabix.bed.gz.dvc | 106.0 B | 6bffd4977ea6d62d96659401cc560769 |
| ENCFF078TXM.tabix.bed.gz.tbi | 106.35 KB | 2f76b5bb98220500a12f9b08da511b79 |
| ENCFF078TXM.tabix.bed.gz.tbi.dvc | 110.0 B | f99872c94d4be330b492052a7f9b2dfe |
| genomic_resource.yaml | 2.57 KB | 1e184ceb953fec6c9fb64ff0f229a0b7 |
| genomic_resource_original.yaml | 2.47 KB | b98687857535681f379d6b19c0894f7e |
| statistics/ |