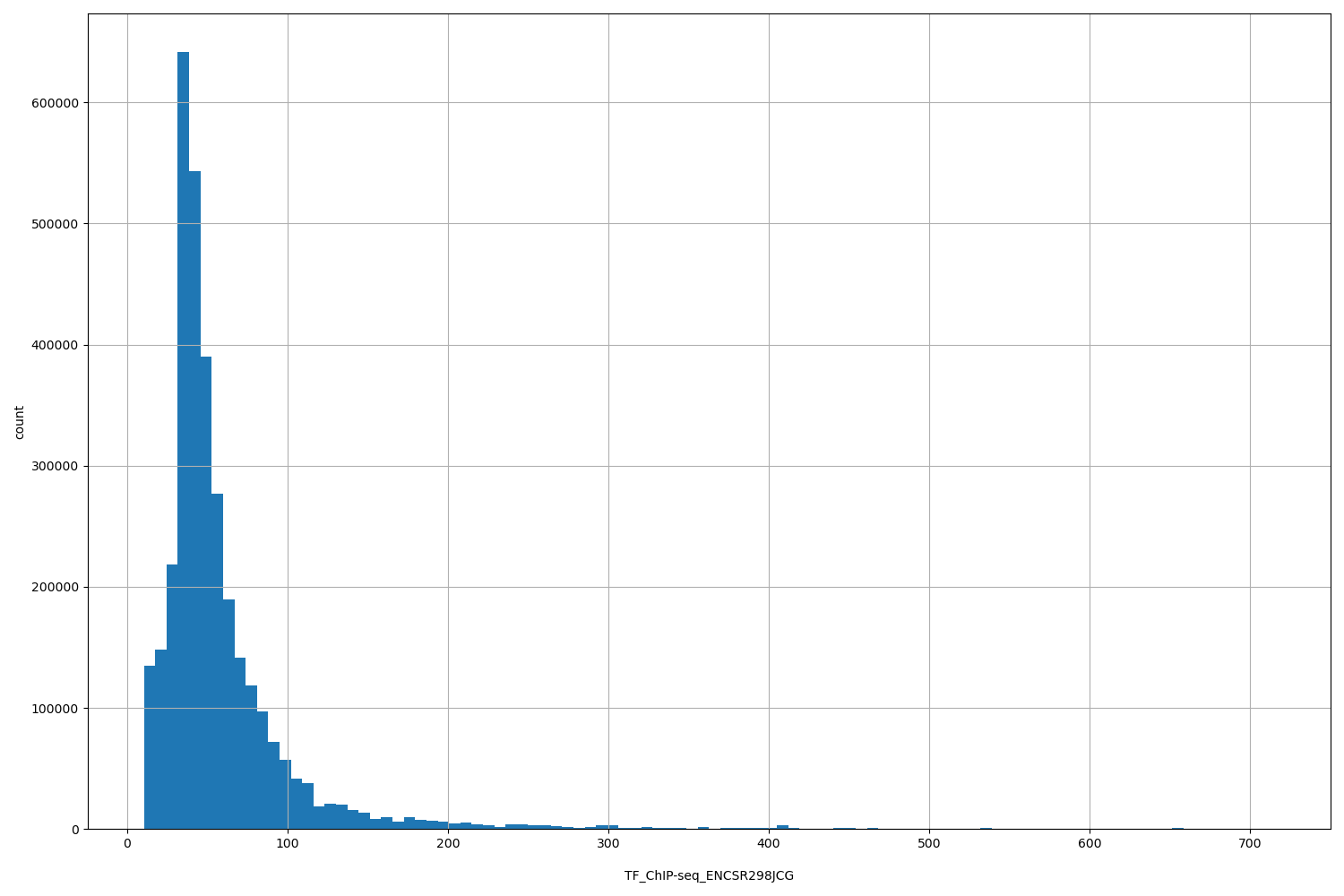
TF_ChIP-seq_ENCSR298JCG
| Id: | TF_ChIP-seq/ENCSR298JCG |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR298JCG [biosamplesummary="Homo sapiens K562" and target="NCOR1"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN723QXU|/analyses/ENCAN723QXU/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_internal_action: File {ENCFF638IIC|/files/ENCFF638IIC/} with status 'released' is derived from file {ENCFF419RSJ|/files/ENCFF419RSJ/} with status 'archived'. audit_warning: Processed alignments file {ENCFF247FGI|/files/ENCFF247FGI/} processed by ChIP-seq ENCODE3 GRCh38 pipeline has 18688822 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting NCOR1-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR298JCG | float |
TF_ChIP-seq_ENCSR298JCG |
TF_ChIP-seq ENCSR298JCG [biosample_summary="Homo sapiens K562" and target="NCOR1"]
|
 |
[10.4, 715] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF638IIC.bed.gz | 164.94 KB | a6fb144af17e1bf26776ea364011337f |
| ENCFF638IIC.bed.gz.dvc | 100.0 B | 7c8bab235a0d752524040462246fffea |
| ENCFF638IIC.tabix.bed.gz | 116.56 KB | 274cb9d1a8846387bd73ffae6abf52e0 |
| ENCFF638IIC.tabix.bed.gz.dvc | 106.0 B | f4d515bd5cdc82c9350a92db6b537a2d |
| ENCFF638IIC.tabix.bed.gz.tbi | 65.89 KB | f34c63fdcab7906104eaba8b261f7722 |
| ENCFF638IIC.tabix.bed.gz.tbi.dvc | 109.0 B | cb359883e6f226c762dbd5ab2b624536 |
| genomic_resource.yaml | 2.02 KB | c2759c18defd44bcb28da1fe0688eb67 |
| genomic_resource_original.yaml | 1.93 KB | 704072714372fc99fec741d275e385d5 |
| statistics/ |