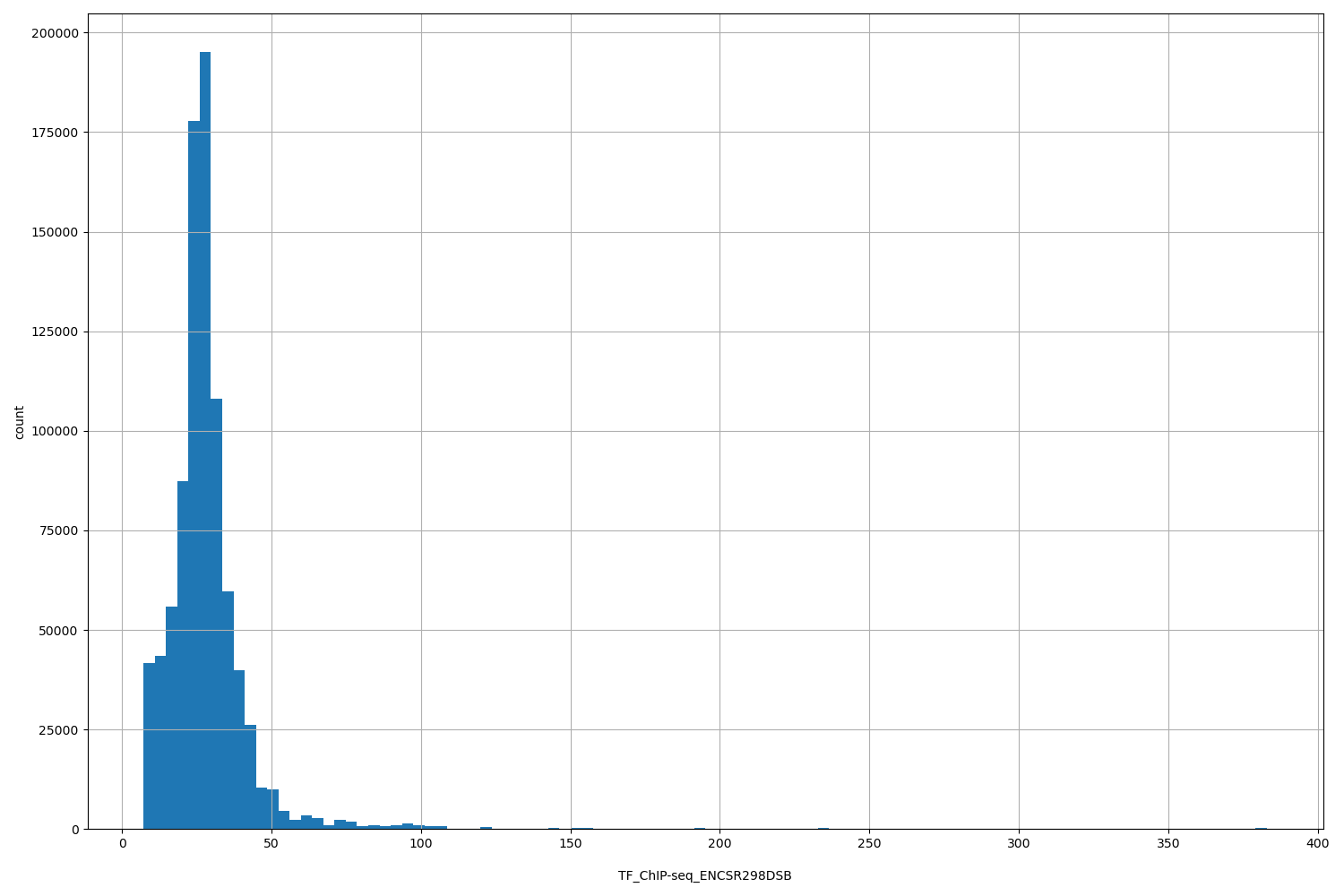
TF_ChIP-seq_ENCSR298DSB
| Id: | TF_ChIP-seq/ENCSR298DSB |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR298DSB [biosamplesummary="Homo sapiens HepG2 genetically modified (insertion) using CRISPR targeting H. sapiens ZNF180" and target="ZNF180"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: genetically modified (insertion) using CRISPR targeting H. sapiens ZNF180 output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN413LBX|/analyses/ENCAN413LBX/} has in progress subobject document {6665f89b-94ed-4e38-a76e-c1cc8cfedfa5|/documents/6665f89b-94ed-4e38-a76e-c1cc8cfedfa5/} audit_internal_action: Released analysis {ENCAN413LBX|/analyses/ENCAN413LBX/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_warning: Processed alignments file {ENCFF066PQO|/files/ENCFF066PQO/} processed by ChIP-seq ENCODE4 v1.4.0 GRCh38 pipeline has 18343821 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting ZNF180-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF405YXO|/files/ENCFF405YXO/} processed by ChIP-seq ENCODE4 v1.4.0 GRCh38 pipeline have a rescue ratio of 1.56 and a self consistency ratio of 3.98. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF263XZK|/files/ENCFF263XZK/} processed by ChIP-seq ENCODE4 v1.4.0 GRCh38 pipeline have a rescue ratio of 1.56 and a self consistency ratio of 3.98. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR298DSB | float |
TF_ChIP-seq_ENCSR298DSB |
TF_ChIP-seq ENCSR298DSB [biosample_summary="Homo sapiens HepG2 genetically modified (insertion) using CRISPR targeting H. sapiens ZNF180" and target="ZNF180"]
|
 |
[7.24, 383] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF405YXO.bed.gz | 52.7 KB | 974ff9cb784a46240da39218a3684fff |
| ENCFF405YXO.bed.gz.dvc | 99.0 B | 73cc6ebfb8b07ebb01c6f3bacad3b137 |
| ENCFF405YXO.tabix.bed.gz | 36.62 KB | 82ae5b562cc5ce063ec77f743ace31fb |
| ENCFF405YXO.tabix.bed.gz.dvc | 105.0 B | 34dd958db442e3d62c2fdd408e752667 |
| ENCFF405YXO.tabix.bed.gz.tbi | 26.85 KB | 56569120458ceda99ad0c1ba8d447ccd |
| ENCFF405YXO.tabix.bed.gz.tbi.dvc | 109.0 B | 65ad90e8ee7f374c46485b7aa8c12d7e |
| genomic_resource.yaml | 3.45 KB | a39b73aec818da273140f4154e6a2386 |
| genomic_resource_original.yaml | 3.28 KB | 5d1568bf5f10faf70be17c4587d6cc7b |
| statistics/ |