TF_ChIP-seq_ENCSR295BIP
| Id: | TF_ChIP-seq/ENCSR295BIP |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR295BIP [biosamplesummary="Homo sapiens HEK293 genetically modified (insertion) using site-specific recombination targeting H. sapiens ZNF777" and target="ZNF777"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: genetically modified (insertion) using site-specific recombination targeting H. sapiens ZNF777 output_type: optimal IDR thresholded peaks audit_internal_action: File {ENCFF804ZLF|/files/ENCFF804ZLF/} with status 'released' is derived from file {ENCFF419RSJ|/files/ENCFF419RSJ/} with status 'archived'. audit_internal_action: Archived analysis {ENCAN667TTZ|/analyses/ENCAN667TTZ/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF221OEX|/files/ENCFF221OEX/}, {ENCFF804ZLF|/files/ENCFF804ZLF/} processed by ChIP-seq ENCODE3 GRCh38 pipeline have a rescue ratio of 1.69 and a self consistency ratio of 3.75. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF720WAE|/files/ENCFF720WAE/}, {ENCFF163VTM|/files/ENCFF163VTM/} processed by ChIP-seq ENCODE3 GRCh38 pipeline have a rescue ratio of 1.69 and a self consistency ratio of 3.75. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
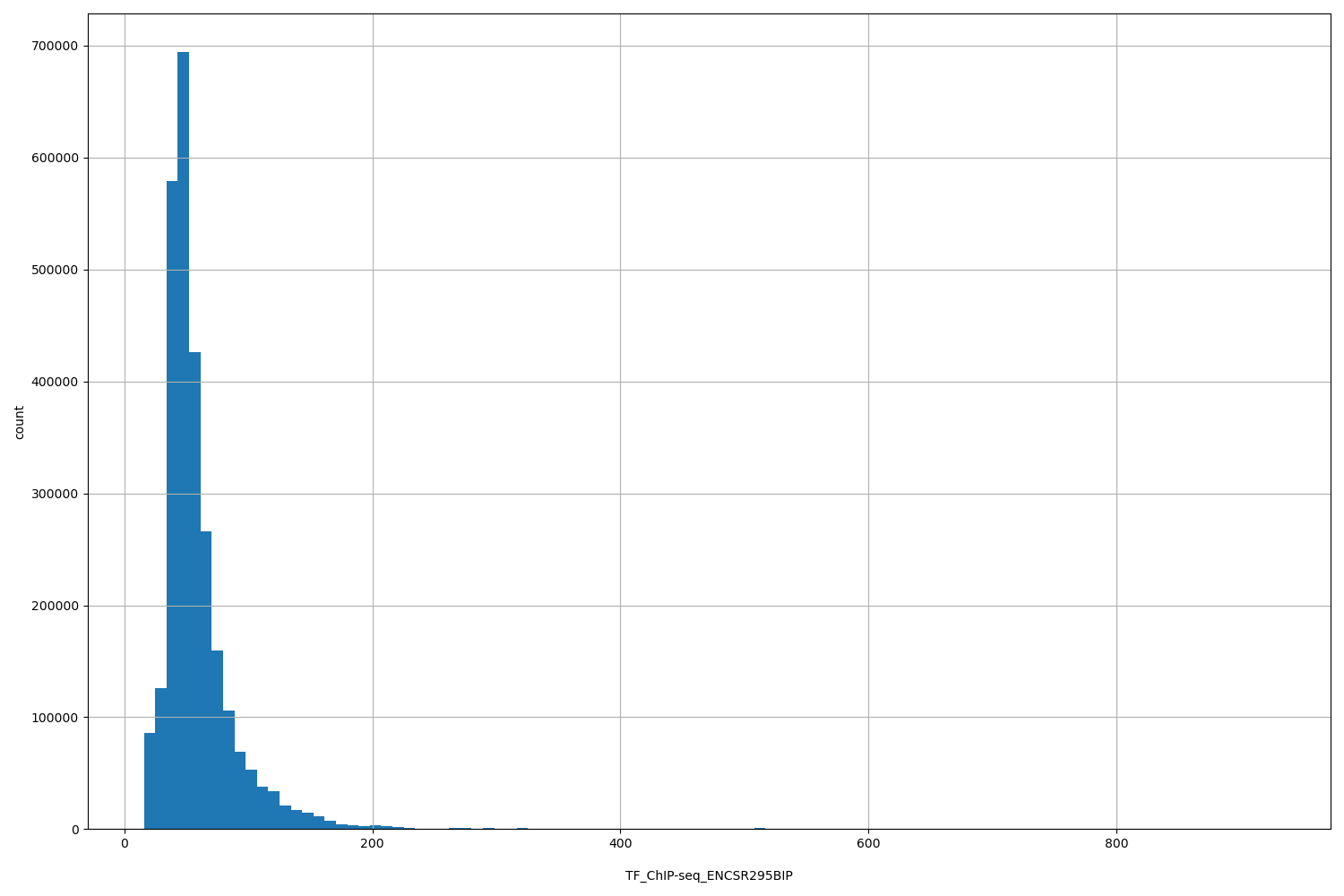
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR295BIP | float |
TF_ChIP-seq_ENCSR295BIP |
TF_ChIP-seq ENCSR295BIP [biosample_summary="Homo sapiens HEK293 genetically modified (insertion) using site-specific recombination targeting H. sapiens ZNF777" and target="ZNF777"]
|
 |
[15.8, 927] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF804ZLF.bed.gz | 142.8 KB | 8eeababfab257e430c4b9aa1f511e66a |
| ENCFF804ZLF.bed.gz.dvc | 100.0 B | c456036e860423f20dea3756a44f55bd |
| ENCFF804ZLF.tabix.bed.gz | 98.64 KB | 8c5fac8937c6a37112758cf1b22e6696 |
| ENCFF804ZLF.tabix.bed.gz.dvc | 106.0 B | 4fb3328cf45265e888845cdebc2644fb |
| ENCFF804ZLF.tabix.bed.gz.tbi | 62.8 KB | 32fd41164a20f50083c481440e7ed4cc |
| ENCFF804ZLF.tabix.bed.gz.tbi.dvc | 109.0 B | ef4f2406a2b76202d5553a386caedfb3 |
| genomic_resource.yaml | 3.1 KB | b096e481674b71dd97d65cb2d2690b02 |
| genomic_resource_original.yaml | 2.92 KB | b3f5e712c9f6ac9a555d77d9b570d285 |
| statistics/ |