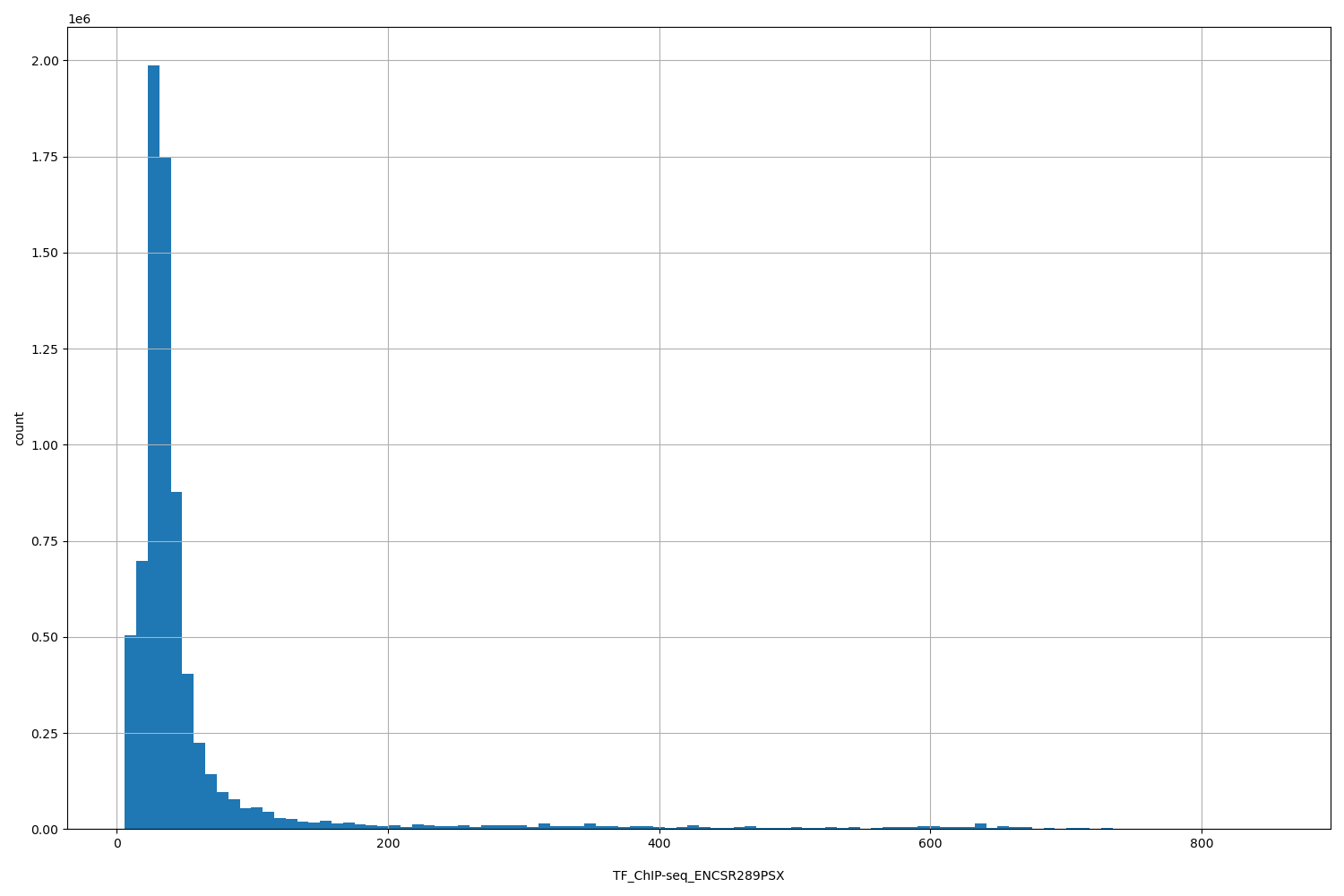
TF_ChIP-seq_ENCSR289PSX
| Id: | TF_ChIP-seq/ENCSR289PSX |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR289PSX [biosamplesummary="Homo sapiens HepG2 genetically modified (insertion) using CRISPR targeting H. sapiens PRDM10" and target="PRDM10"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: genetically modified (insertion) using CRISPR targeting H. sapiens PRDM10 output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN914RAR|/analyses/ENCAN914RAR/} has in progress subobject document {6f435fd3-3884-42d4-8907-09d09c134754|/documents/6f435fd3-3884-42d4-8907-09d09c134754/} audit_internal_action: Released analysis {ENCAN914RAR|/analyses/ENCAN914RAR/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_warning: Processed alignments file {ENCFF190USS|/files/ENCFF190USS/} processed by ChIP-seq ENCODE4 v1.4.0 GRCh38 pipeline has 18618935 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting PRDM10-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Processed alignments file {ENCFF968NER|/files/ENCFF968NER/} processed by ChIP-seq ENCODE4 v1.4.0 GRCh38 pipeline has 16670483 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting PRDM10-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR289PSX | float |
TF_ChIP-seq_ENCSR289PSX |
TF_ChIP-seq ENCSR289PSX [biosample_summary="Homo sapiens HepG2 genetically modified (insertion) using CRISPR targeting H. sapiens PRDM10" and target="PRDM10"]
|
 |
[6.07, 853] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF668GIP.bed.gz | 310.4 KB | e0121439c3b639e0d85b60b26c395065 |
| ENCFF668GIP.bed.gz.dvc | 100.0 B | 30fa878fe7f831887a118ee6ee5950d7 |
| ENCFF668GIP.tabix.bed.gz | 232.67 KB | 08ec5ebf12388e77285b20fd60cec945 |
| ENCFF668GIP.tabix.bed.gz.dvc | 106.0 B | c6714684fa1713d58299f6a3b81546e9 |
| ENCFF668GIP.tabix.bed.gz.tbi | 105.87 KB | dfc050e098544bebc009ae616e66ca08 |
| ENCFF668GIP.tabix.bed.gz.tbi.dvc | 110.0 B | 544addac2bc9cbbfcac69fff033c1c82 |
| genomic_resource.yaml | 2.89 KB | 214b5ebeb9a8fbaf5c53b562bbd076da |
| genomic_resource_original.yaml | 2.72 KB | f3675fcb1a06e6c61208ea1b313e0f53 |
| statistics/ |