TF_ChIP-seq_ENCSR251OVJ
| Id: | TF_ChIP-seq/ENCSR251OVJ |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR251OVJ [biosamplesummary="Homo sapiens GM12878" and target="SMAD5"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: optimal IDR thresholded peaks audit_internal_action: File {ENCFF855SJG|/files/ENCFF855SJG/} with status 'released' is derived from file {ENCFF419RSJ|/files/ENCFF419RSJ/} with status 'archived'. audit_internal_action: Archived analysis {ENCAN835DJR|/analyses/ENCAN835DJR/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Processed alignments file {ENCFF346SPJ|/files/ENCFF346SPJ/} processed by ChIP-seq ENCODE3 GRCh38 pipeline has 16707955 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting SMAD5-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
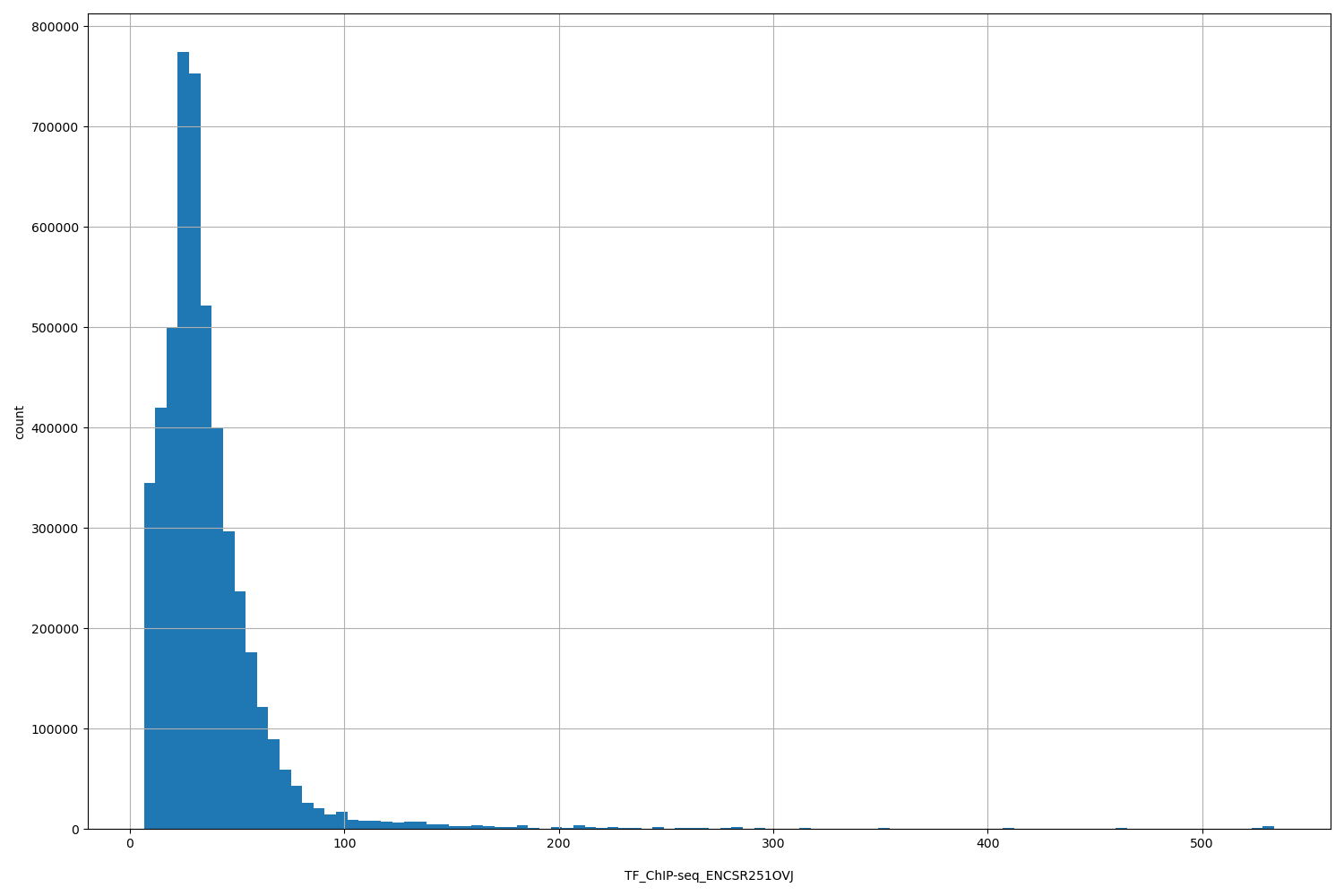
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR251OVJ | float |
TF_ChIP-seq_ENCSR251OVJ |
TF_ChIP-seq ENCSR251OVJ [biosample_summary="Homo sapiens GM12878" and target="SMAD5"]
|
 |
[6.53, 534] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF855SJG.bed.gz | 208.65 KB | 3a660a224b00c18129ab276553653d1f |
| ENCFF855SJG.bed.gz.dvc | 100.0 B | 0b7f7f809619e9373045b946e50e8506 |
| ENCFF855SJG.tabix.bed.gz | 168.03 KB | 0fc7c75397be1a4ec7da70ca416ba1ee |
| ENCFF855SJG.tabix.bed.gz.dvc | 106.0 B | 345b00bcffdebaa90ab3db2ab2270f36 |
| ENCFF855SJG.tabix.bed.gz.tbi | 69.8 KB | 69253af75081068d73f577e2221cf8c6 |
| ENCFF855SJG.tabix.bed.gz.tbi.dvc | 109.0 B | 5a966c4ed75b053cf33a869eb6056acd |
| genomic_resource.yaml | 2.03 KB | cefa69bfa3419cb998a5f95290e31092 |
| genomic_resource_original.yaml | 1.93 KB | 2655d0029ffddab03f76dc5044e21e3d |
| statistics/ |