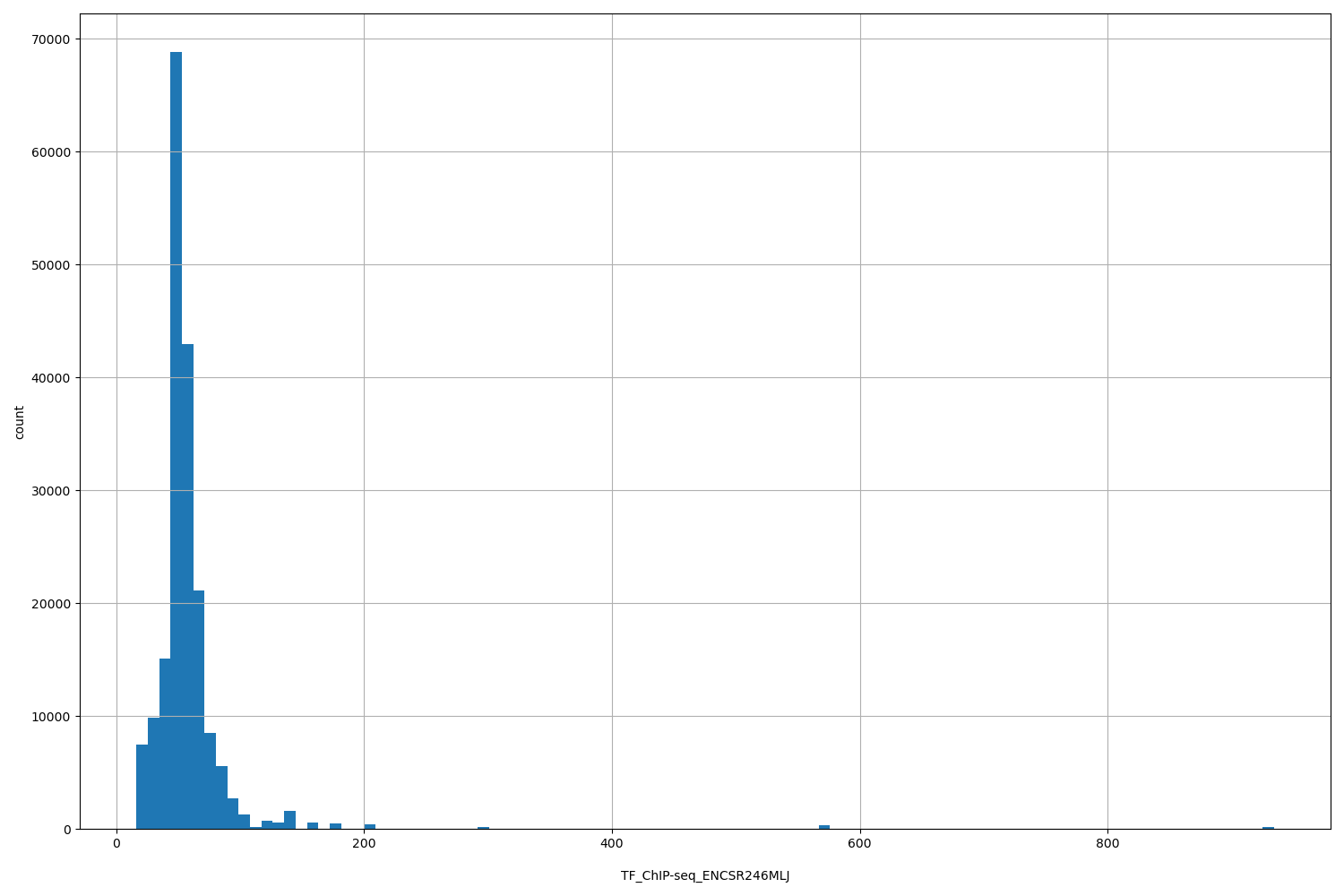
TF_ChIP-seq_ENCSR246MLJ
| Id: | TF_ChIP-seq/ENCSR246MLJ |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR246MLJ [biosamplesummary="Homo sapiens 22Rv1" and target="ZFX"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: optimal IDR thresholded peaks audit_internal_action: File {ENCFF447UZC|/files/ENCFF447UZC/} with status 'released' is derived from file {ENCFF419RSJ|/files/ENCFF419RSJ/} with status 'archived'. audit_internal_action: Archived analysis {ENCAN937JJR|/analyses/ENCAN937JJR/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF550TZM|/files/ENCFF550TZM/}, {ENCFF552UMS|/files/ENCFF552UMS/} processed by ChIP-seq ENCODE3 GRCh38 pipeline have a rescue ratio of 1.85 and a self consistency ratio of 2.12. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF056PDQ|/files/ENCFF056PDQ/}, {ENCFF447UZC|/files/ENCFF447UZC/} processed by ChIP-seq ENCODE3 GRCh38 pipeline have a rescue ratio of 1.85 and a self consistency ratio of 2.12. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR246MLJ | float |
TF_ChIP-seq_ENCSR246MLJ |
TF_ChIP-seq ENCSR246MLJ [biosample_summary="Homo sapiens 22Rv1" and target="ZFX"]
|
 |
[16.1, 934] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF447UZC.bed.gz | 7.34 KB | 4f39c6fcd46ab6d0794e047f87e03de7 |
| ENCFF447UZC.bed.gz.dvc | 98.0 B | 1bc5f26658a741091bdfbfb13a0ec868 |
| ENCFF447UZC.tabix.bed.gz | 5.34 KB | baa6a3f2ab7a468fd81d310d4b920d85 |
| ENCFF447UZC.tabix.bed.gz.dvc | 104.0 B | 855122a252d6849f89d6d401ffc32f75 |
| ENCFF447UZC.tabix.bed.gz.tbi | 7.72 KB | 5560f0227bfea8404d540a1a850355f8 |
| ENCFF447UZC.tabix.bed.gz.tbi.dvc | 108.0 B | 68d6049ef1d3c1912b12d986f61a6e95 |
| genomic_resource.yaml | 2.66 KB | 0ad90898f32d5ae35da6cccbbe0dfc3c |
| genomic_resource_original.yaml | 2.56 KB | 59c35a621ef2969ce77d15fafd5e1290 |
| statistics/ |