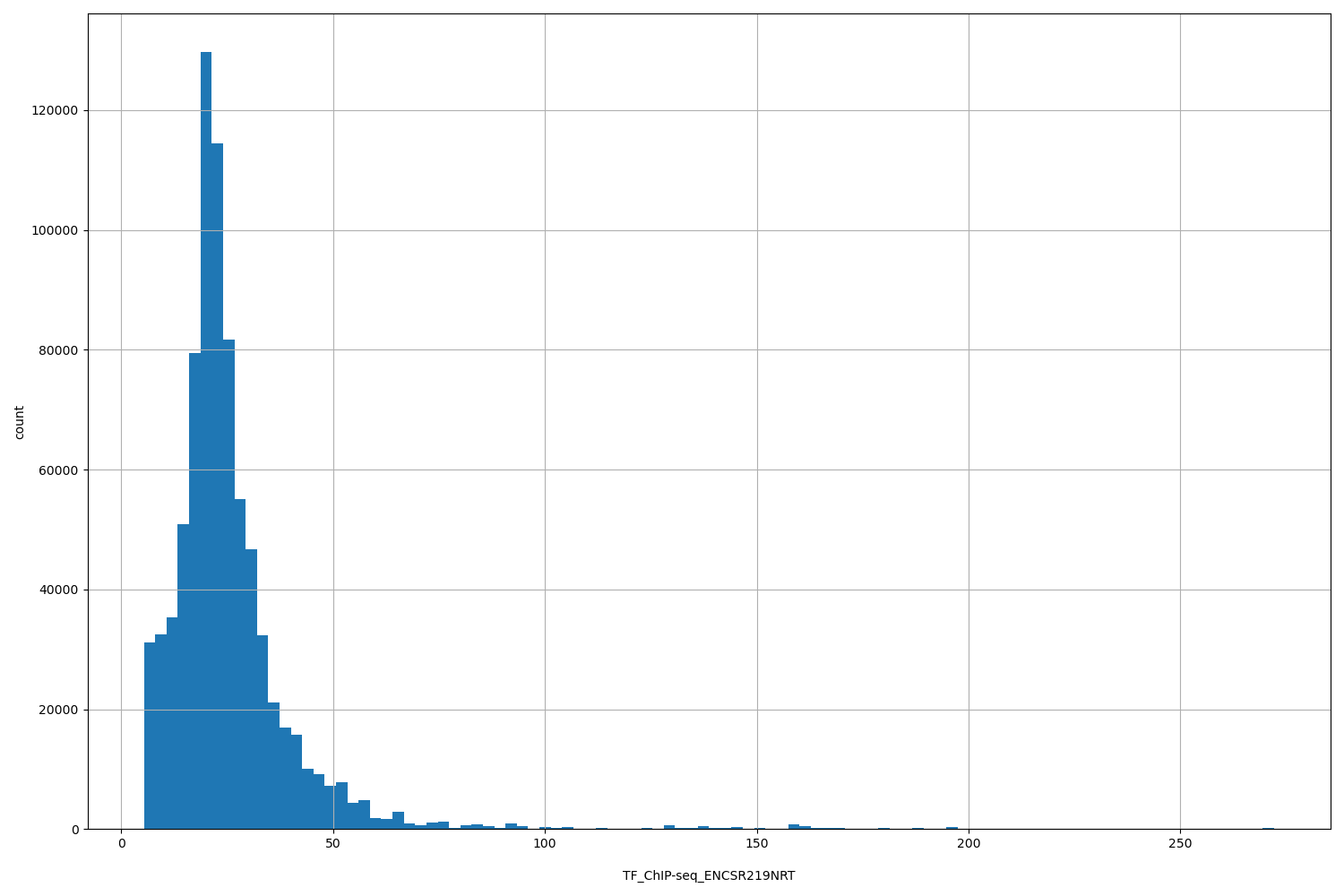
TF_ChIP-seq_ENCSR219NRT
| Id: | TF_ChIP-seq/ENCSR219NRT |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR219NRT [biosamplesummary="Homo sapiens HepG2" and target="GTF2F1"] |
| Description: |
status: released biological_replicates: Rep 2, Rep 3 summary: output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN747FSC|/analyses/ENCAN747FSC/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Processed alignments file {ENCFF204YCT|/files/ENCFF204YCT/} processed by ChIP-seq ENCODE3 hg19 pipeline has 19028365 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting GTF2F1-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR219NRT | float |
TF_ChIP-seq_ENCSR219NRT |
TF_ChIP-seq ENCSR219NRT [biosample_summary="Homo sapiens HepG2" and target="GTF2F1"]
|
 |
[5.33, 272] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF694UJI.bed.gz | 47.41 KB | 8c61a3021ba1648497bcc6c7e1a1d4d7 |
| ENCFF694UJI.bed.gz.dvc | 99.0 B | 3b3a0383bd2d70c2728c271dbf3b6c12 |
| ENCFF694UJI.tabix.bed.gz | 34.13 KB | 12a4cae496afc3e69dd2fae677c4606d |
| ENCFF694UJI.tabix.bed.gz.dvc | 105.0 B | 1a29a8d5ade6ea983922108db30e870d |
| ENCFF694UJI.tabix.bed.gz.tbi | 27.28 KB | b165305afc98433d1a5579b7574431d0 |
| ENCFF694UJI.tabix.bed.gz.tbi.dvc | 109.0 B | 4b2ea754d8ab0e42a133a051957a9da0 |
| genomic_resource.yaml | 1.84 KB | 9da5ca8abfd74660ad6c0eb5b888bca6 |
| genomic_resource_original.yaml | 1.75 KB | 04343821d79748d284ba1601c3c79394 |
| statistics/ |