TF_ChIP-seq_ENCSR207PFI
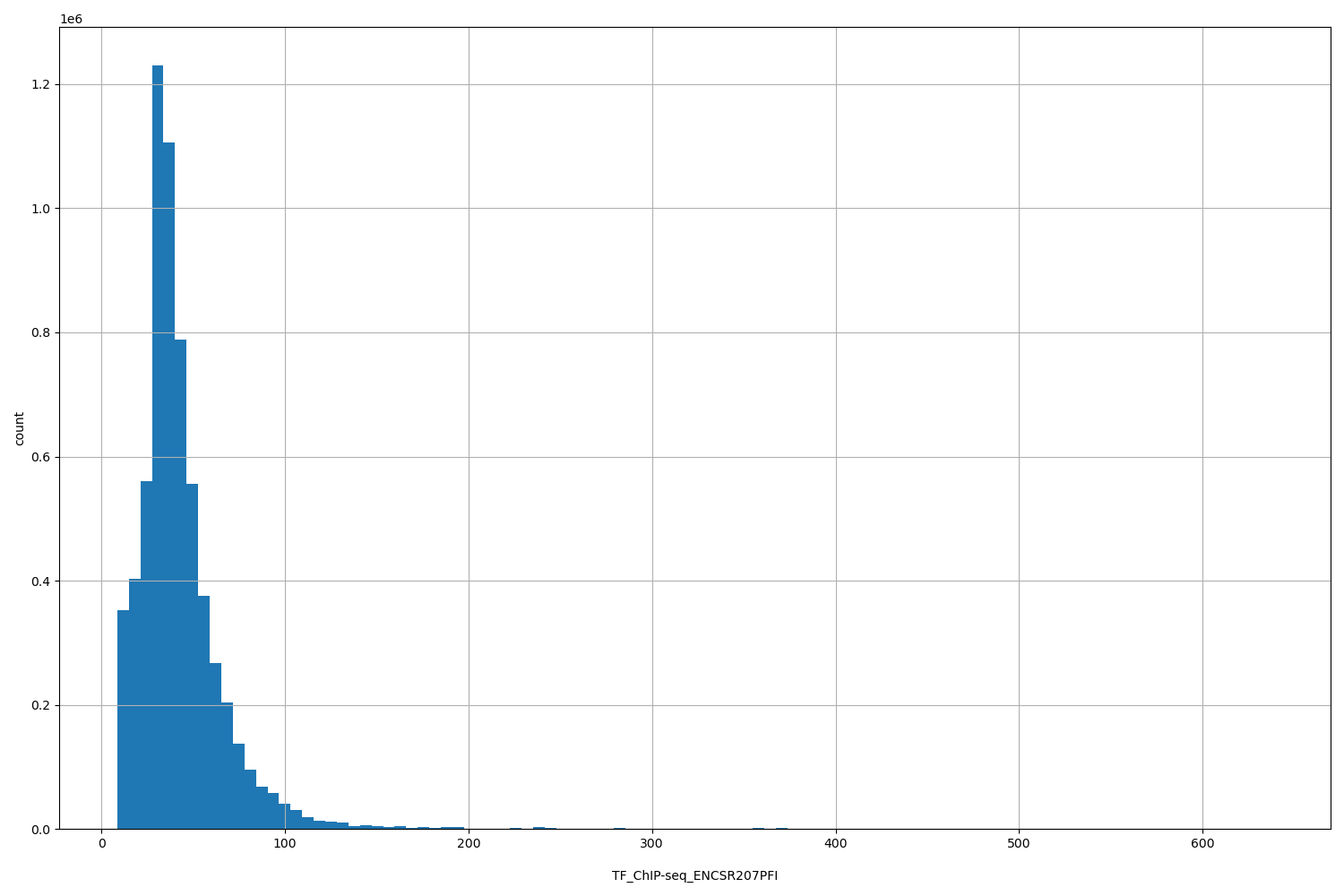
| Id: | TF_ChIP-seq/ENCSR207PFI |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR207PFI [biosamplesummary="Homo sapiens GM12878" and target="ZBED1"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN307ZVL|/analyses/ENCAN307ZVL/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_internal_action: File {ENCFF630FLK|/files/ENCFF630FLK/} with status 'released' is derived from file {ENCFF419RSJ|/files/ENCFF419RSJ/} with status 'archived'. audit_internal_action: Released file {ENCFF630FLK|/files/ENCFF630FLK/} has replaced subobject file {07fa8e17-071b-4208-94b0-77b31daf9f01|/files/07fa8e17-071b-4208-94b0-77b31daf9f01/} audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF630FLK|/files/ENCFF630FLK/}, {ENCFF907JXH|/files/ENCFF907JXH/} processed by ChIP-seq ENCODE3 GRCh38 pipeline have a rescue ratio of 1.53 and a self consistency ratio of 3.00. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF688FED|/files/ENCFF688FED/}, {ENCFF125JVH|/files/ENCFF125JVH/} processed by ChIP-seq ENCODE3 GRCh38 pipeline have a rescue ratio of 1.53 and a self consistency ratio of 3.00. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR207PFI | float |
TF_ChIP-seq_ENCSR207PFI |
TF_ChIP-seq ENCSR207PFI [biosample_summary="Homo sapiens GM12878" and target="ZBED1"]
|
 |
[8.6, 638] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF630FLK.bed.gz | 258.2 KB | 1307e25bd4c8b822ca0dca5bf395edec |
| ENCFF630FLK.bed.gz.dvc | 100.0 B | 14839248f9332850528f1fb5cbaeaa0f |
| ENCFF630FLK.tabix.bed.gz | 194.7 KB | a94efcc7bca870453ae0c22922d5bb37 |
| ENCFF630FLK.tabix.bed.gz.dvc | 106.0 B | 9b8d6129c766c005e25efaa6fea372f5 |
| ENCFF630FLK.tabix.bed.gz.tbi | 86.57 KB | 23fe75e8098d72725a47c9a080adf8ba |
| ENCFF630FLK.tabix.bed.gz.tbi.dvc | 109.0 B | fa8c939b88abdb618a9fdbfba264893e |
| genomic_resource.yaml | 2.87 KB | 55a688771a9c34b0c9bd798fd0eb3320 |
| genomic_resource_original.yaml | 2.77 KB | 7ee88933701215c37aeb9df34c72f189 |
| statistics/ |