TF_ChIP-seq_ENCSR200CUA
| Id: | TF_ChIP-seq/ENCSR200CUA |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR200CUA [biosamplesummary="Homo sapiens MCF-7" and target="HDGF"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN375WKE|/analyses/ENCAN375WKE/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF758RLH|/files/ENCFF758RLH/}, {ENCFF445ANP|/files/ENCFF445ANP/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 2.16 and a self consistency ratio of 1.16. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF020JAD|/files/ENCFF020JAD/}, {ENCFF176YMC|/files/ENCFF176YMC/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 2.16 and a self consistency ratio of 1.16. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
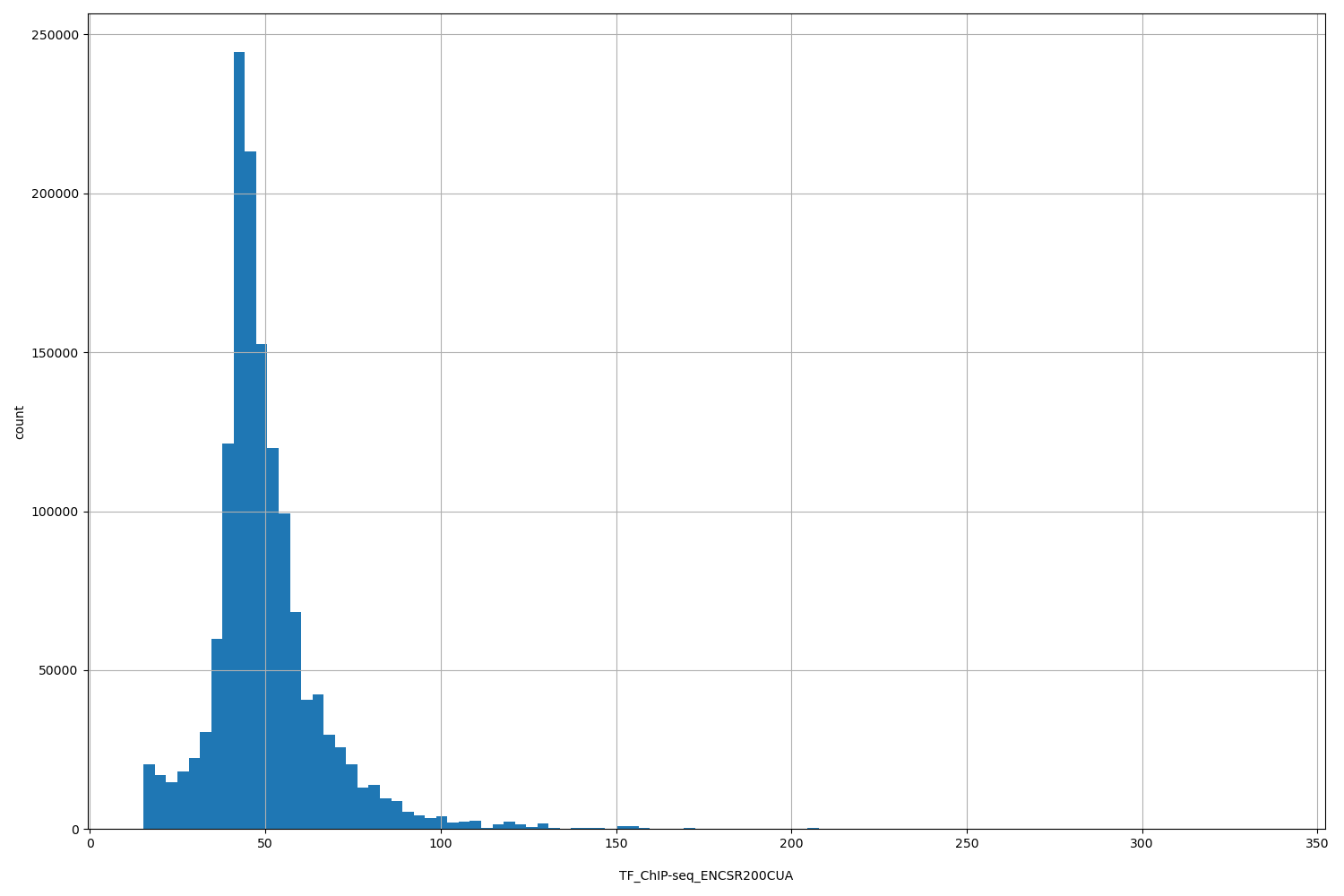
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR200CUA | float |
TF_ChIP-seq_ENCSR200CUA |
TF_ChIP-seq ENCSR200CUA [biosample_summary="Homo sapiens MCF-7" and target="HDGF"]
|
 |
[15.3, 336] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF445ANP.bed.gz | 76.57 KB | 667ac466c60ad43c4538ca0023c227e1 |
| ENCFF445ANP.bed.gz.dvc | 99.0 B | b20d67de3c142c9e5a2f2b8d1fdf4fd5 |
| ENCFF445ANP.tabix.bed.gz | 51.04 KB | 3a7c9e22725bb8dfb40b265cb0ed6795 |
| ENCFF445ANP.tabix.bed.gz.dvc | 105.0 B | c390bc9a3d9671f2abb8b0c6902de19e |
| ENCFF445ANP.tabix.bed.gz.tbi | 31.71 KB | aedd0cc1a223cbae81450409f8467570 |
| ENCFF445ANP.tabix.bed.gz.tbi.dvc | 109.0 B | 4cf659fadc5bff0644702ae052c4b160 |
| genomic_resource.yaml | 2.47 KB | d96b64527a1d7d0469a1834d7132a7b5 |
| genomic_resource_original.yaml | 2.38 KB | 12cfd52cfb36141711c308a9f66b088c |
| statistics/ |