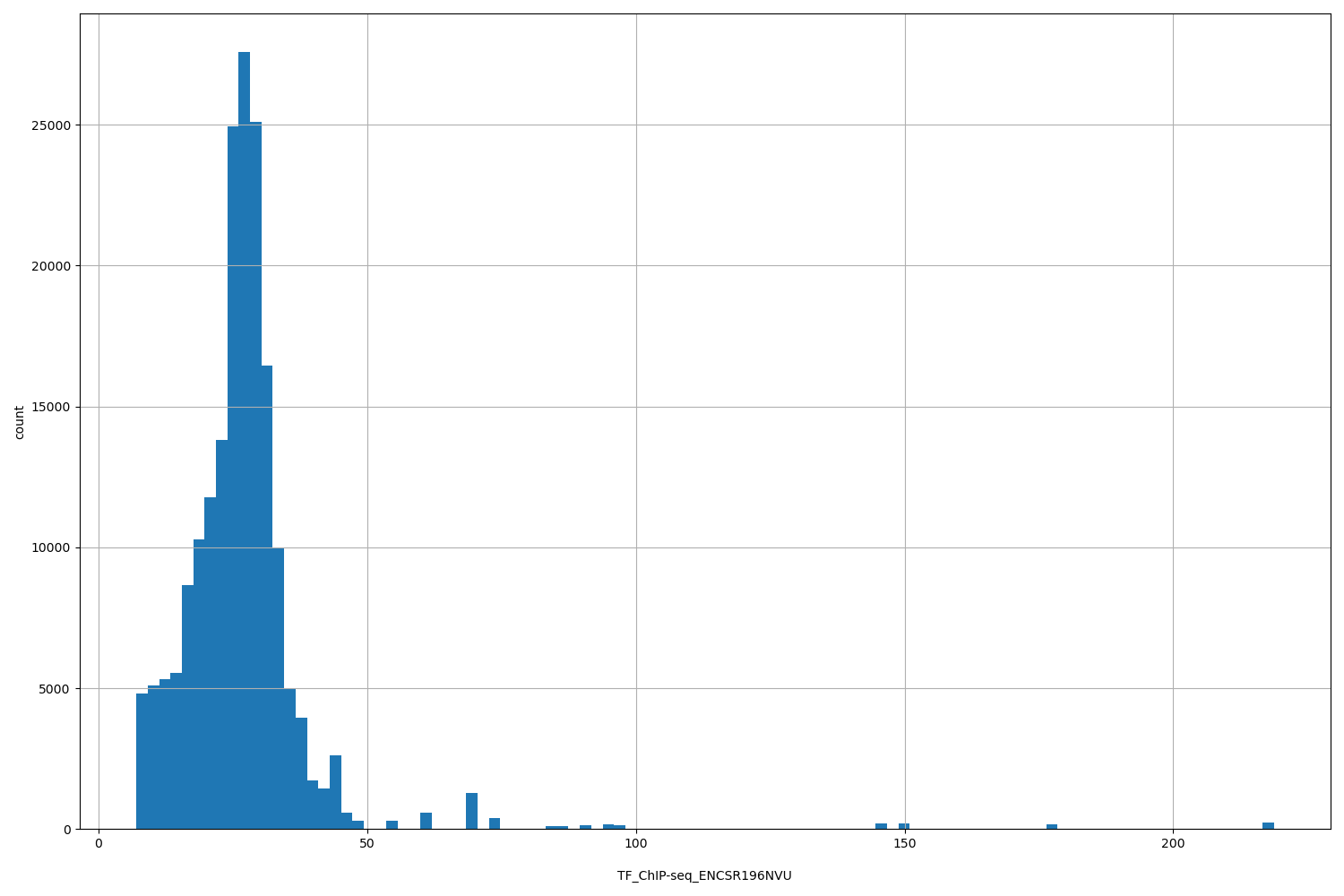
TF_ChIP-seq_ENCSR196NVU
| Id: | TF_ChIP-seq/ENCSR196NVU |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR196NVU [biosamplesummary="Homo sapiens HepG2 genetically modified (insertion) using CRISPR targeting H. sapiens ATRX" and target="ATRX"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: genetically modified (insertion) using CRISPR targeting H. sapiens ATRX output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN902YZL|/analyses/ENCAN902YZL/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_internal_action: Released analysis {ENCAN902YZL|/analyses/ENCAN902YZL/} has in progress subobject document {e79e415b-ed2d-420b-95a7-f9be1bdb34e4|/documents/e79e415b-ed2d-420b-95a7-f9be1bdb34e4/} audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF998FLA|/files/ENCFF998FLA/} processed by ChIP-seq ENCODE4 v1.4.0 GRCh38 pipeline have a rescue ratio of 1.70 and a self consistency ratio of 5.82. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF195YNC|/files/ENCFF195YNC/} processed by ChIP-seq ENCODE4 v1.4.0 GRCh38 pipeline have a rescue ratio of 1.70 and a self consistency ratio of 5.82. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR196NVU | float |
TF_ChIP-seq_ENCSR196NVU |
TF_ChIP-seq ENCSR196NVU [biosample_summary="Homo sapiens HepG2 genetically modified (insertion) using CRISPR targeting H. sapiens ATRX" and target="ATRX"]
|
 |
[7.04, 219] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF998FLA.bed.gz | 12.53 KB | 3e77ccaaed739c7b859cea1eede39e51 |
| ENCFF998FLA.bed.gz.dvc | 99.0 B | 35a90760f7dfd0fedc506a62cdf52ade |
| ENCFF998FLA.tabix.bed.gz | 8.74 KB | 5140c502ba8ebf5d3340217cc9243236 |
| ENCFF998FLA.tabix.bed.gz.dvc | 104.0 B | fe2acd3448351ef72e2e2709f2f9b9ab |
| ENCFF998FLA.tabix.bed.gz.tbi | 9.98 KB | 59ea6cb67691b4bee9a74bcdfa192f9a |
| ENCFF998FLA.tabix.bed.gz.tbi.dvc | 109.0 B | 3500b5b197878e060c2e61ca62686d01 |
| genomic_resource.yaml | 2.94 KB | 8211f29cd5ed6c1df9ea3a17c65d6d99 |
| genomic_resource_original.yaml | 2.78 KB | 7f61eb078b057d9971611a4a6925bf2c |
| statistics/ |