TF_ChIP-seq_ENCSR178QVJ
| Id: | TF_ChIP-seq/ENCSR178QVJ |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR178QVJ [biosamplesummary="Homo sapiens HEK293 genetically modified (insertion) using site-specific recombination targeting H. sapiens ZNF274" and target="ZNF274"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: genetically modified (insertion) using site-specific recombination targeting H. sapiens ZNF274 output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN724DTT|/analyses/ENCAN724DTT/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF265HWY|/files/ENCFF265HWY/}, {ENCFF027RLD|/files/ENCFF027RLD/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 2.18 and a self consistency ratio of 1.79. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF128MHV|/files/ENCFF128MHV/}, {ENCFF297VSS|/files/ENCFF297VSS/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 2.18 and a self consistency ratio of 1.79. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
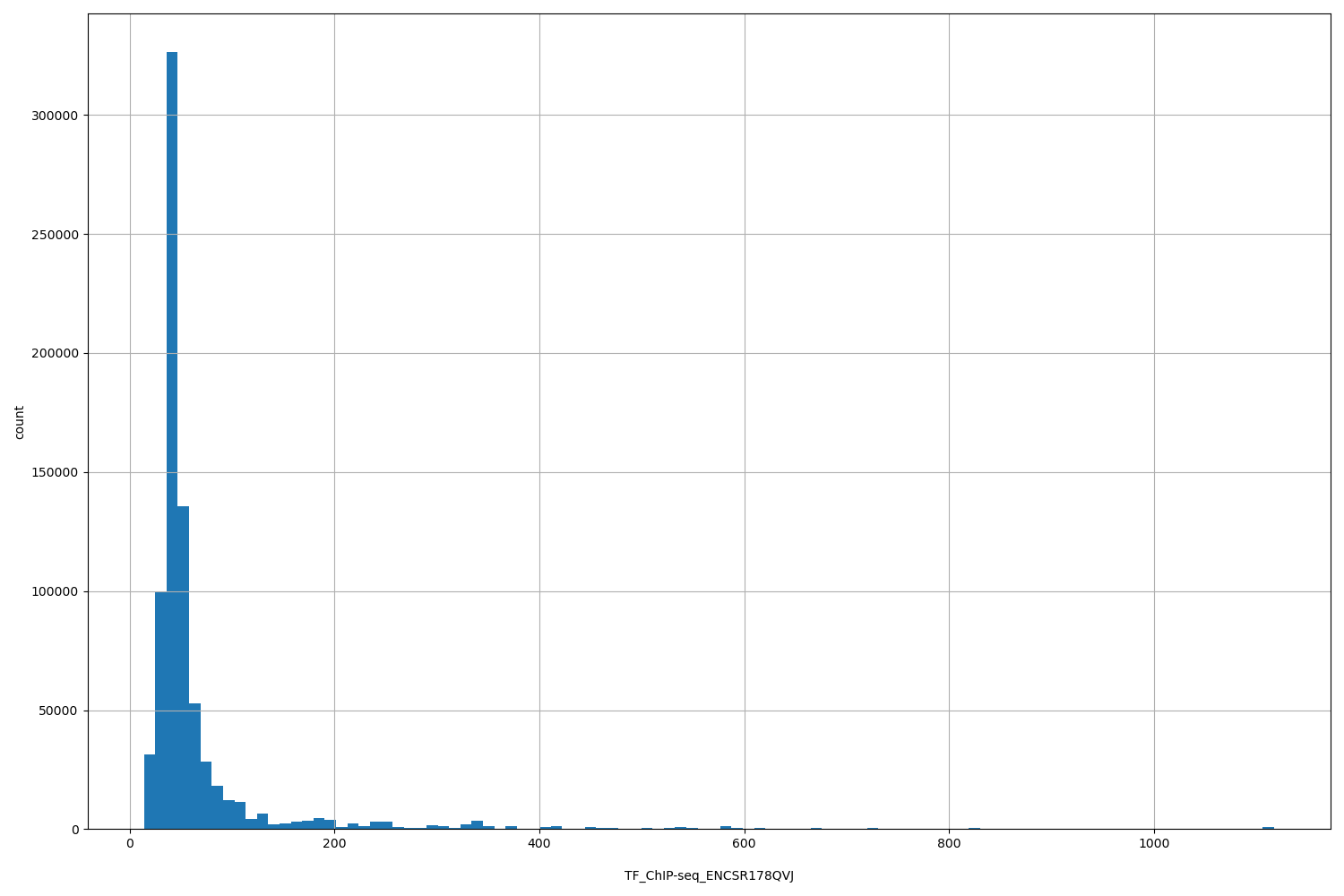
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR178QVJ | float |
TF_ChIP-seq_ENCSR178QVJ |
TF_ChIP-seq ENCSR178QVJ [biosample_summary="Homo sapiens HEK293 genetically modified (insertion) using site-specific recombination targeting H. sapiens ZNF274" and target="ZNF274"]
|
 |
[13.9, 1.12e+03] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF265HWY.bed.gz | 44.81 KB | 1747130c7d70af102753a38155223689 |
| ENCFF265HWY.bed.gz.dvc | 99.0 B | d0796a735f55d9c57668aeb166c2337f |
| ENCFF265HWY.tabix.bed.gz | 29.7 KB | 66979a15e652987f99a811e8df3c0acd |
| ENCFF265HWY.tabix.bed.gz.dvc | 105.0 B | da1c51a4d777c754b074ef3f564897a7 |
| ENCFF265HWY.tabix.bed.gz.tbi | 24.67 KB | 84c740356a4a840ff72c6200114f940d |
| ENCFF265HWY.tabix.bed.gz.tbi.dvc | 109.0 B | a8769454f8d98987cdbb473ba78a6011 |
| genomic_resource.yaml | 2.91 KB | d5bcc54e25484324747a3510c333fc6f |
| genomic_resource_original.yaml | 2.73 KB | adabcbbb2c2dc2f9375a41cc1db215f2 |
| statistics/ |