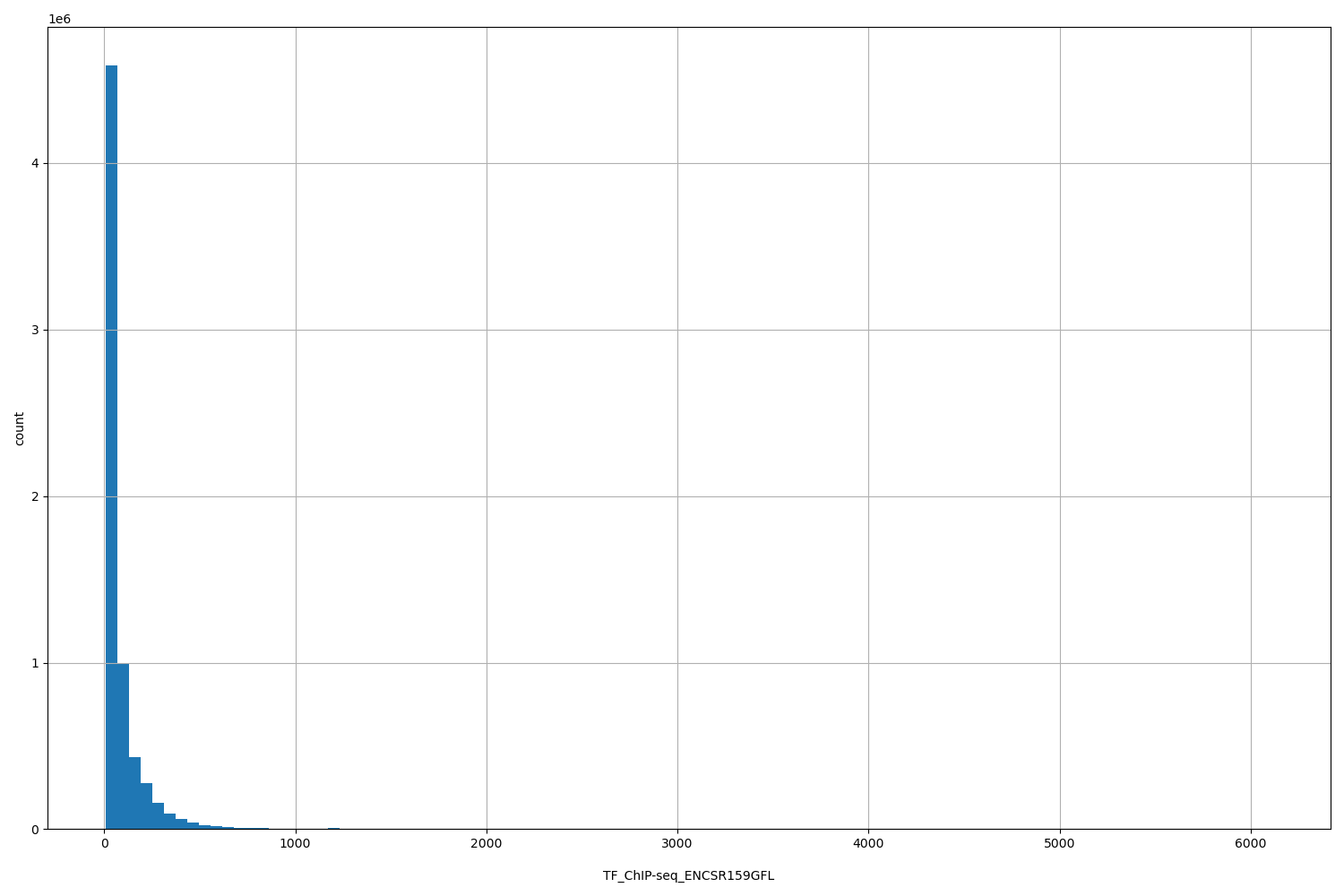
TF_ChIP-seq_ENCSR159GFL
| Id: | TF_ChIP-seq/ENCSR159GFL |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR159GFL [biosamplesummary="Homo sapiens HEK293 genetically modified (insertion) using site-specific recombination targeting H. sapiens ZNF518A" and target="ZNF518A"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: genetically modified (insertion) using site-specific recombination targeting H. sapiens ZNF518A output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN939ISU|/analyses/ENCAN939ISU/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Processed alignments file {ENCFF213TEN|/files/ENCFF213TEN/} processed by ChIP-seq ENCODE3 hg19 pipeline has 18863474 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting ZNF518A-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Processed alignments file {ENCFF224CNS|/files/ENCFF224CNS/} processed by ChIP-seq ENCODE3 hg19 pipeline has 15984375 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting ZNF518A-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR159GFL | float |
TF_ChIP-seq_ENCSR159GFL |
TF_ChIP-seq ENCSR159GFL [biosample_summary="Homo sapiens HEK293 genetically modified (insertion) using site-specific recombination targeting H. sapiens ZNF518A" and target="ZNF518A"]
|
 |
[9.58, 6.11e+03] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF415VBF.bed.gz | 339.95 KB | 4ec7c0243a797e956a9d8b6cffc90c85 |
| ENCFF415VBF.bed.gz.dvc | 100.0 B | da58eb1c423518f7508a171817fd2ac2 |
| ENCFF415VBF.tabix.bed.gz | 241.57 KB | a8189bd6ca8cdd93afb2fc6312903abc |
| ENCFF415VBF.tabix.bed.gz.dvc | 106.0 B | 4875af5c980a773526bf4194b1e609ee |
| ENCFF415VBF.tabix.bed.gz.tbi | 136.63 KB | a1f34679b9a057752050f0b7ac611cab |
| ENCFF415VBF.tabix.bed.gz.tbi.dvc | 110.0 B | 9693257b3b4f440c10c91b47c1c4e210 |
| genomic_resource.yaml | 2.78 KB | 2be7eeee0ab08b2dba60b17d946c82b9 |
| genomic_resource_original.yaml | 2.6 KB | 3ad3fa362e40f06d011be9af631f4bee |
| statistics/ |