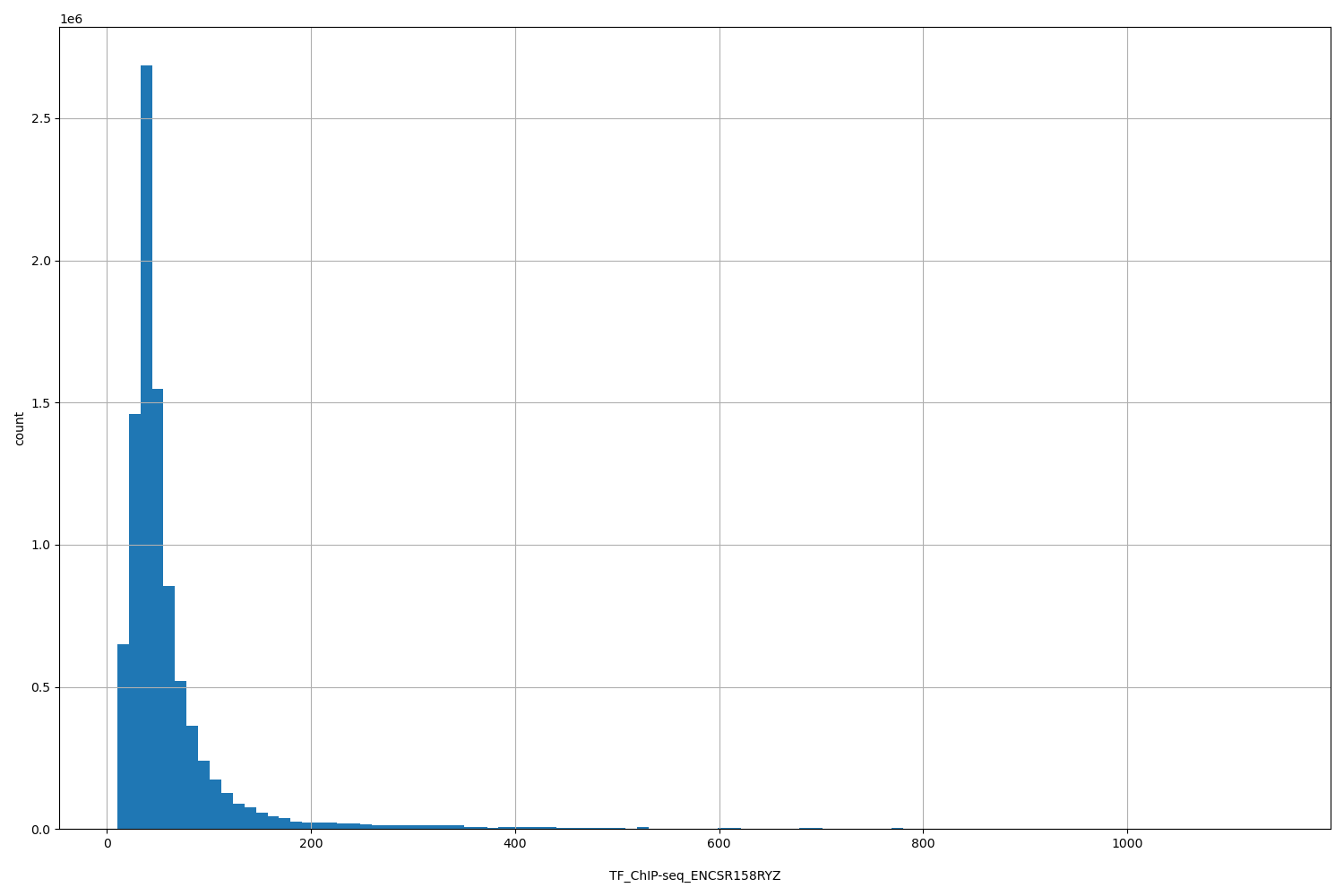
TF_ChIP-seq_ENCSR158RYZ
| Id: | TF_ChIP-seq/ENCSR158RYZ |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR158RYZ [biosamplesummary="Homo sapiens K562 genetically modified (insertion) using CRISPR targeting H. sapiens ZBTB40" and target="ZBTB40"] |
| Description: |
status: archived biological_replicates: Rep 3, Rep 4 summary: genetically modified (insertion) using CRISPR targeting H. sapiens ZBTB40 output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN735JZB|/analyses/ENCAN735JZB/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF413TRG|/files/ENCFF413TRG/}, {ENCFF995IAC|/files/ENCFF995IAC/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 1.36 and a self consistency ratio of 2.35. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF986DSB|/files/ENCFF986DSB/}, {ENCFF841JIB|/files/ENCFF841JIB/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 1.36 and a self consistency ratio of 2.35. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR158RYZ | float |
TF_ChIP-seq_ENCSR158RYZ |
TF_ChIP-seq ENCSR158RYZ [biosample_summary="Homo sapiens K562 genetically modified (insertion) using CRISPR targeting H. sapiens ZBTB40" and target="ZBTB40"]
|
 |
[10.2, 1.14e+03] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF995IAC.bed.gz | 342.17 KB | 5fc18b2a8f2ddbf62a95eb4350681ef0 |
| ENCFF995IAC.bed.gz.dvc | 100.0 B | 277316fcf658c6d258d52da38f026cde |
| ENCFF995IAC.tabix.bed.gz | 259.28 KB | 347879bfed763f7c3959cebbc9ef0f98 |
| ENCFF995IAC.tabix.bed.gz.dvc | 106.0 B | a3f84bcd5a383299d7c7ccd61e0f60d3 |
| ENCFF995IAC.tabix.bed.gz.tbi | 114.3 KB | 64b6aa18e9fc5ed4cc7d8efd4c1bd9b4 |
| ENCFF995IAC.tabix.bed.gz.tbi.dvc | 110.0 B | 97195dee85424841223636e2d17f0adf |
| genomic_resource.yaml | 2.8 KB | e90b57e70161a45ffb71a9aed0d81600 |
| genomic_resource_original.yaml | 2.64 KB | 7f3661bf1352b0b553d62ac0cf74d5e3 |
| statistics/ |