TF_ChIP-seq_ENCSR138YYY
| Id: | TF_ChIP-seq/ENCSR138YYY |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR138YYY [biosamplesummary="Homo sapiens K562" and target="GABPB1"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: optimal IDR thresholded peaks audit_internal_action: File {ENCFF700DXR|/files/ENCFF700DXR/} with status 'released' is derived from file {ENCFF419RSJ|/files/ENCFF419RSJ/} with status 'archived'. audit_internal_action: Released file {ENCFF700DXR|/files/ENCFF700DXR/} has replaced subobject file {b6bdf6b4-1260-431e-9151-275ea0092b7c|/files/b6bdf6b4-1260-431e-9151-275ea0092b7c/} audit_internal_action: Archived analysis {ENCAN071JCD|/analyses/ENCAN071JCD/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: PBC1 (PCR Bottlenecking Coefficient 1, M1/M_distinct) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where some reads map (M_distinct). A PBC1 value in the range 0 - 0.5 is severe bottlenecking, 0.5 - 0.8 is moderate bottlenecking, 0.8 - 0.9 is mild bottlenecking, and > 0.9 is no bottlenecking. PBC1 value > 0.9 is recommended, but > 0.8 is acceptable. ENCODE processed alignments file {ENCFF838GWC|/files/ENCFF838GWC/} processed by ChIP-seq ENCODE3 GRCh38 pipeline was generated from a library with PBC1 value of 0.84. audit_warning: PBC2 (PCR Bottlenecking Coefficient 2, M1/M2) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where two reads map uniquely (M2). A PBC2 value in the range 0 - 1 is severe bottlenecking, 1 - 3 is moderate bottlenecking, 3 - 10 is mild bottlenecking, > 10 is no bottlenecking. PBC2 value > 10 is recommended, but > 3 is acceptable. ENCODE processed alignments file {ENCFF838GWC|/files/ENCFF838GWC/} processed by ChIP-seq ENCODE3 GRCh38 pipeline was generated from a library with PBC2 value of 6.23. audit_warning: PBC1 (PCR Bottlenecking Coefficient 1, M1/M_distinct) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where some reads map (M_distinct). A PBC1 value in the range 0 - 0.5 is severe bottlenecking, 0.5 - 0.8 is moderate bottlenecking, 0.8 - 0.9 is mild bottlenecking, and > 0.9 is no bottlenecking. PBC1 value > 0.9 is recommended, but > 0.8 is acceptable. ENCODE processed alignments file {ENCFF183DVK|/files/ENCFF183DVK/} processed by ChIP-seq ENCODE3 GRCh38 pipeline was generated from a library with PBC1 value of 0.89. audit_warning: PBC2 (PCR Bottlenecking Coefficient 2, M1/M2) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where two reads map uniquely (M2). A PBC2 value in the range 0 - 1 is severe bottlenecking, 1 - 3 is moderate bottlenecking, 3 - 10 is mild bottlenecking, > 10 is no bottlenecking. PBC2 value > 10 is recommended, but > 3 is acceptable. ENCODE processed alignments file {ENCFF183DVK|/files/ENCFF183DVK/} processed by ChIP-seq ENCODE3 GRCh38 pipeline was generated from a library with PBC2 value of 8.90. |
| Labels: |
|
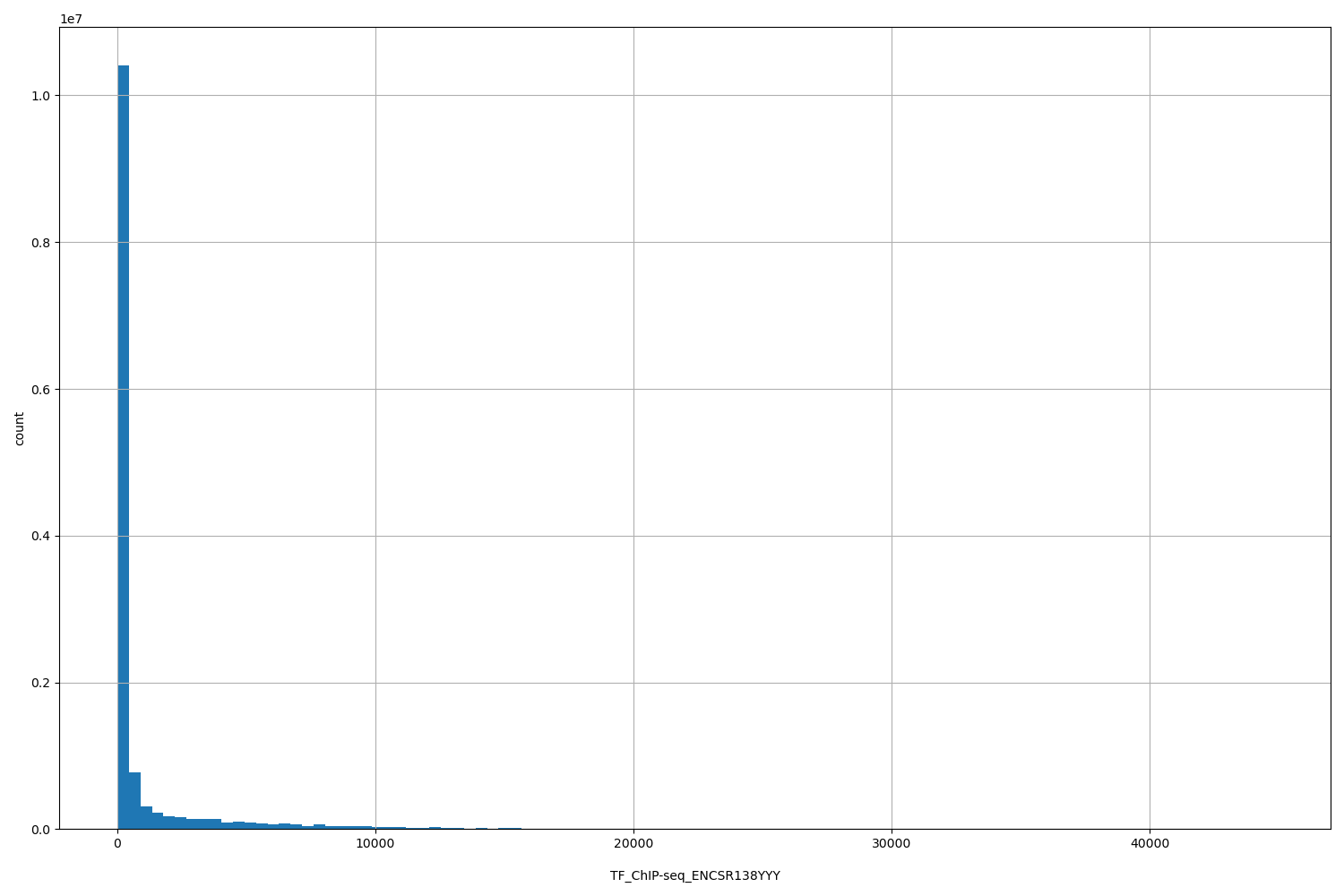
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR138YYY | float |
TF_ChIP-seq_ENCSR138YYY |
TF_ChIP-seq ENCSR138YYY [biosample_summary="Homo sapiens K562" and target="GABPB1"]
|
 |
[10.8, 4.48e+04] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF700DXR.bed.gz | 543.92 KB | 3762f1d9d39f122dad2f23e26972d833 |
| ENCFF700DXR.bed.gz.dvc | 100.0 B | d519cf37d0c023195ffa323e2f93a6b8 |
| ENCFF700DXR.tabix.bed.gz | 373.68 KB | d9a87d23b1f0001cd55fc60bd22dc4aa |
| ENCFF700DXR.tabix.bed.gz.dvc | 106.0 B | 9759fdebc5ebc57d3e87d9657f36c41d |
| ENCFF700DXR.tabix.bed.gz.tbi | 167.09 KB | 2c4330deda1fc2888f1300e55c201306 |
| ENCFF700DXR.tabix.bed.gz.tbi.dvc | 110.0 B | 844240bc1172b21b3a84777e4aa8cbe1 |
| genomic_resource.yaml | 4.88 KB | 8408869c1e43b341fc6b3e259894883e |
| genomic_resource_original.yaml | 4.78 KB | ff042e1eecdb293d8728ebb95c466ca2 |
| statistics/ |