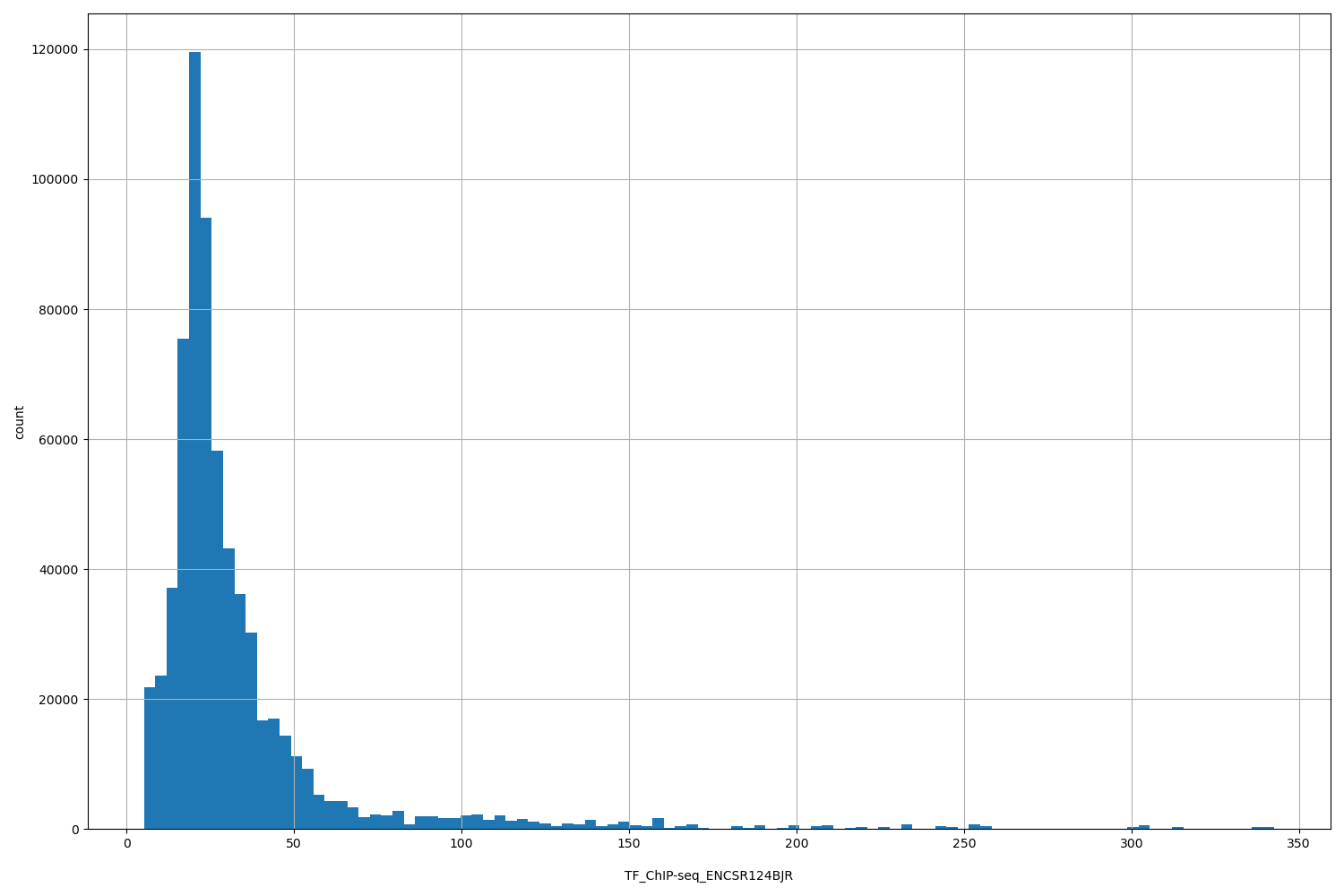
TF_ChIP-seq_ENCSR124BJR
| Id: | TF_ChIP-seq/ENCSR124BJR |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR124BJR [biosamplesummary="Homo sapiens K562" and target="ETV6"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN131DKH|/analyses/ENCAN131DKH/} has in progress subobject document {e11c306e-46d4-4e41-847e-78f5134da02b|/documents/e11c306e-46d4-4e41-847e-78f5134da02b/} audit_internal_action: Released analysis {ENCAN131DKH|/analyses/ENCAN131DKH/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_warning: Processed alignments file {ENCFF366IFJ|/files/ENCFF366IFJ/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 19331550 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting ETV6-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF584QFY|/files/ENCFF584QFY/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline have a rescue ratio of 1.65 and a self consistency ratio of 5.23. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR124BJR | float |
TF_ChIP-seq_ENCSR124BJR |
TF_ChIP-seq ENCSR124BJR [biosample_summary="Homo sapiens K562" and target="ETV6"]
|
 |
[5.14, 343] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF584QFY.bed.gz | 45.28 KB | d5eedcafc74f619ebd8af31068b52ae1 |
| ENCFF584QFY.bed.gz.dvc | 99.0 B | e85982ca0c4fae6d9e23741d4bd339ca |
| ENCFF584QFY.tabix.bed.gz | 31.22 KB | d8e76e0d9d69bcb01c1e05acf93a9baa |
| ENCFF584QFY.tabix.bed.gz.dvc | 105.0 B | a7290f94fb5e63319e44e306adcd6b4f |
| ENCFF584QFY.tabix.bed.gz.tbi | 26.3 KB | e36e2b45c76206a492055ddd483c4fae |
| ENCFF584QFY.tabix.bed.gz.tbi.dvc | 109.0 B | 5f53c3d7b534d1b14cc5271a1453d6ba |
| genomic_resource.yaml | 2.6 KB | 381938a742f4edc82a3986a8827a72df |
| genomic_resource_original.yaml | 2.5 KB | d7d860dd154dea0887b35b0d9ed50ffb |
| statistics/ |