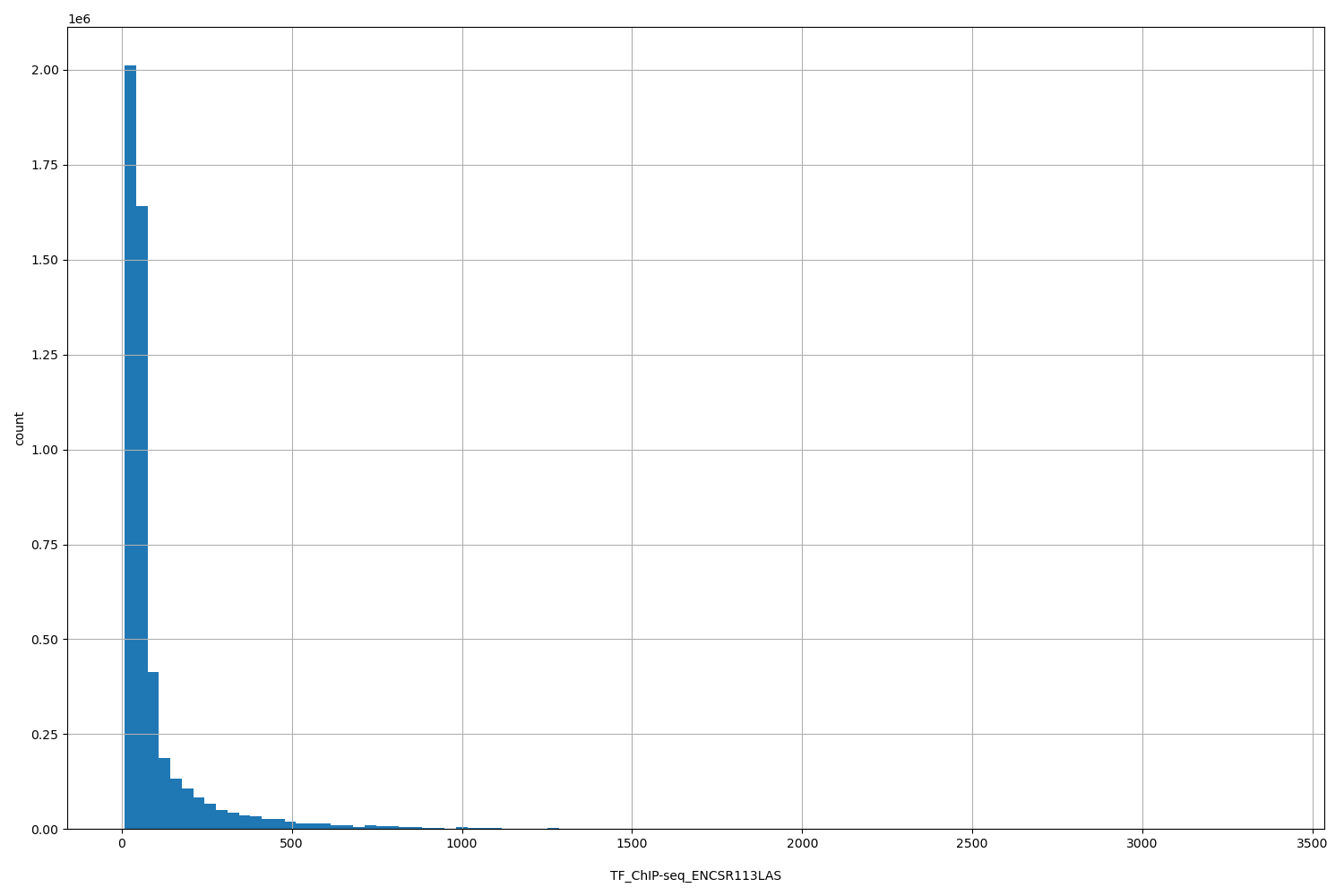
TF_ChIP-seq_ENCSR113LAS
| Id: | TF_ChIP-seq/ENCSR113LAS |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR113LAS [biosamplesummary="Homo sapiens K562" and target="MTA2"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: optimal IDR thresholded peaks audit_internal_action: File {ENCFF713ZVD|/files/ENCFF713ZVD/} with status 'released' is derived from file {ENCFF419RSJ|/files/ENCFF419RSJ/} with status 'archived'. audit_internal_action: Archived analysis {ENCAN810VVR|/analyses/ENCAN810VVR/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Processed alignments file {ENCFF254IUH|/files/ENCFF254IUH/} processed by ChIP-seq ENCODE3 GRCh38 pipeline has 15996994 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting MTA2-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Processed alignments file {ENCFF167NVK|/files/ENCFF167NVK/} processed by ChIP-seq ENCODE3 GRCh38 pipeline has 15608553 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting MTA2-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR113LAS | float |
TF_ChIP-seq_ENCSR113LAS |
TF_ChIP-seq ENCSR113LAS [biosample_summary="Homo sapiens K562" and target="MTA2"]
|
 |
[8.69, 3.37e+03] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF713ZVD.bed.gz | 304.14 KB | 7d5c422e2e0fbd38738e7f2e5ba6653d |
| ENCFF713ZVD.bed.gz.dvc | 100.0 B | b1819e10d66d37cce29470a95a6944fd |
| ENCFF713ZVD.tabix.bed.gz | 205.94 KB | c0e815911ba600889eda3d4e38f3991b |
| ENCFF713ZVD.tabix.bed.gz.dvc | 106.0 B | 849e971c1858d7ddedf6dc324b1c0550 |
| ENCFF713ZVD.tabix.bed.gz.tbi | 117.33 KB | e642b4ccb3a075398ebc20fb10f93307 |
| ENCFF713ZVD.tabix.bed.gz.tbi.dvc | 110.0 B | 77b78eda333f1448d8e959648d966a94 |
| genomic_resource.yaml | 2.51 KB | 02a4bab67b5157088f8e788a8c86cbe3 |
| genomic_resource_original.yaml | 2.42 KB | e0c55cdab98f678167c9c2a861f76991 |
| statistics/ |