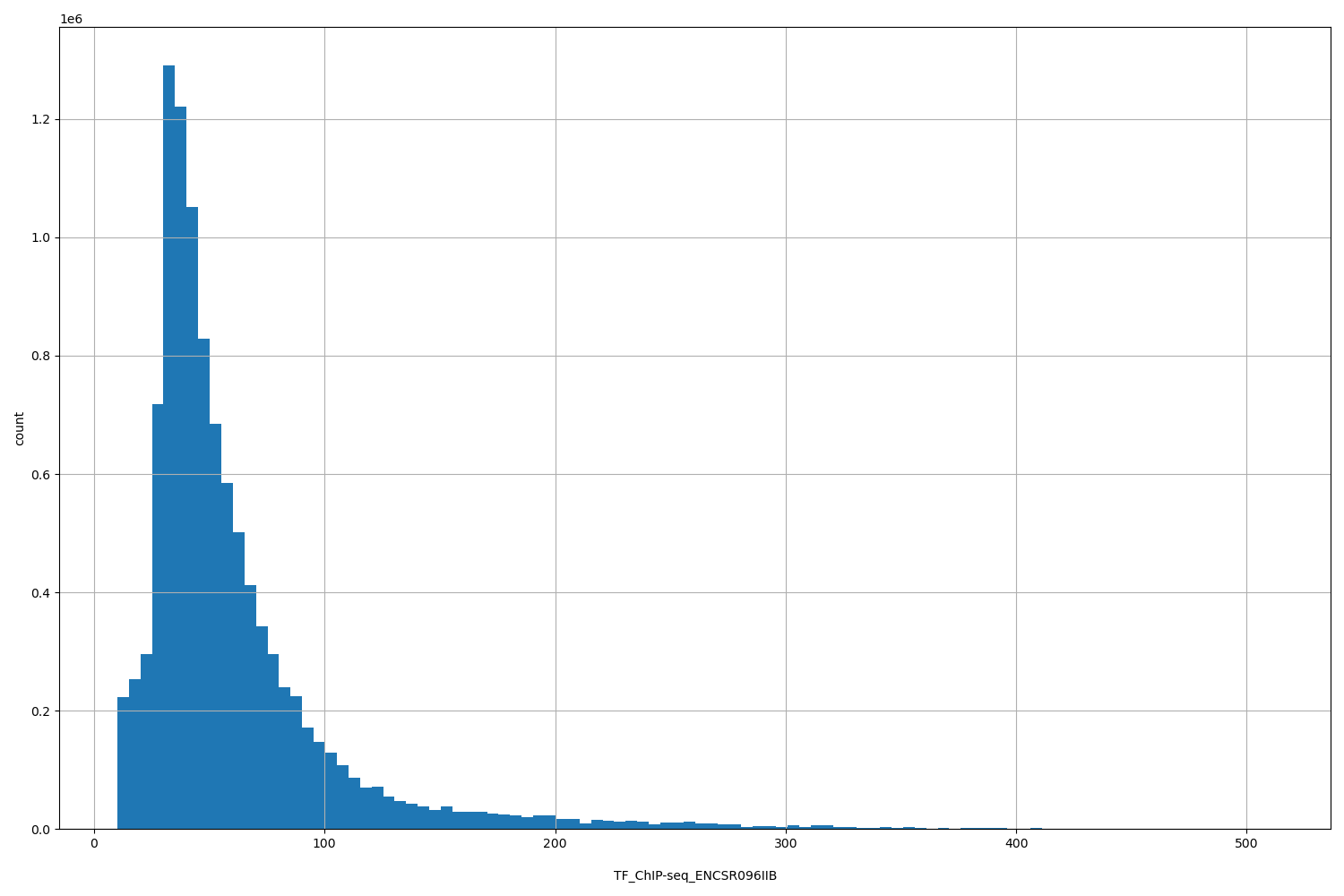
TF_ChIP-seq_ENCSR096IIB
| Id: | TF_ChIP-seq/ENCSR096IIB |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR096IIB [biosamplesummary="Homo sapiens HepG2 genetically modified (insertion) using CRISPR targeting H. sapiens ETV5" and target="ETV5"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: genetically modified (insertion) using CRISPR targeting H. sapiens ETV5 output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN808EUA|/analyses/ENCAN808EUA/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF539NNX|/files/ENCFF539NNX/}, {ENCFF060HVP|/files/ENCFF060HVP/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 1.65 and a self consistency ratio of 3.35. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF860ZQP|/files/ENCFF860ZQP/}, {ENCFF864YJE|/files/ENCFF864YJE/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 1.65 and a self consistency ratio of 3.35. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR096IIB | float |
TF_ChIP-seq_ENCSR096IIB |
TF_ChIP-seq ENCSR096IIB [biosample_summary="Homo sapiens HepG2 genetically modified (insertion) using CRISPR targeting H. sapiens ETV5" and target="ETV5"]
|
 |
[10.1, 511] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF864YJE.bed.gz | 519.36 KB | a952095c309e04251a39ad29756b5fc1 |
| ENCFF864YJE.bed.gz.dvc | 100.0 B | c5e34eae22fa85cb75dd610def9c1fb1 |
| ENCFF864YJE.tabix.bed.gz | 404.08 KB | 463936630b2273f1281528982ad87773 |
| ENCFF864YJE.tabix.bed.gz.dvc | 106.0 B | 53b4e4e38db3ccc8b38e888629e7c48f |
| ENCFF864YJE.tabix.bed.gz.tbi | 170.87 KB | 5d92fe3f65f9e6ec0e4ba54062392c80 |
| ENCFF864YJE.tabix.bed.gz.tbi.dvc | 110.0 B | 7c378cac3a904ac52e78b546ff580dbe |
| genomic_resource.yaml | 2.79 KB | 338be2afe221be8665612db86bf6bc7c |
| genomic_resource_original.yaml | 2.63 KB | a104120b9b0e87b4badeff44e1fa2c71 |
| statistics/ |