TF_ChIP-seq_ENCSR081WLS
| Id: | TF_ChIP-seq/ENCSR081WLS |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR081WLS [biosamplesummary="Homo sapiens HepG2" and target="RAD51"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN040GRI|/analyses/ENCAN040GRI/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF840MPB|/files/ENCFF840MPB/}, {ENCFF031VTR|/files/ENCFF031VTR/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 1.95 and a self consistency ratio of 2.66. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF353OUV|/files/ENCFF353OUV/}, {ENCFF681NJT|/files/ENCFF681NJT/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 1.95 and a self consistency ratio of 2.66. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
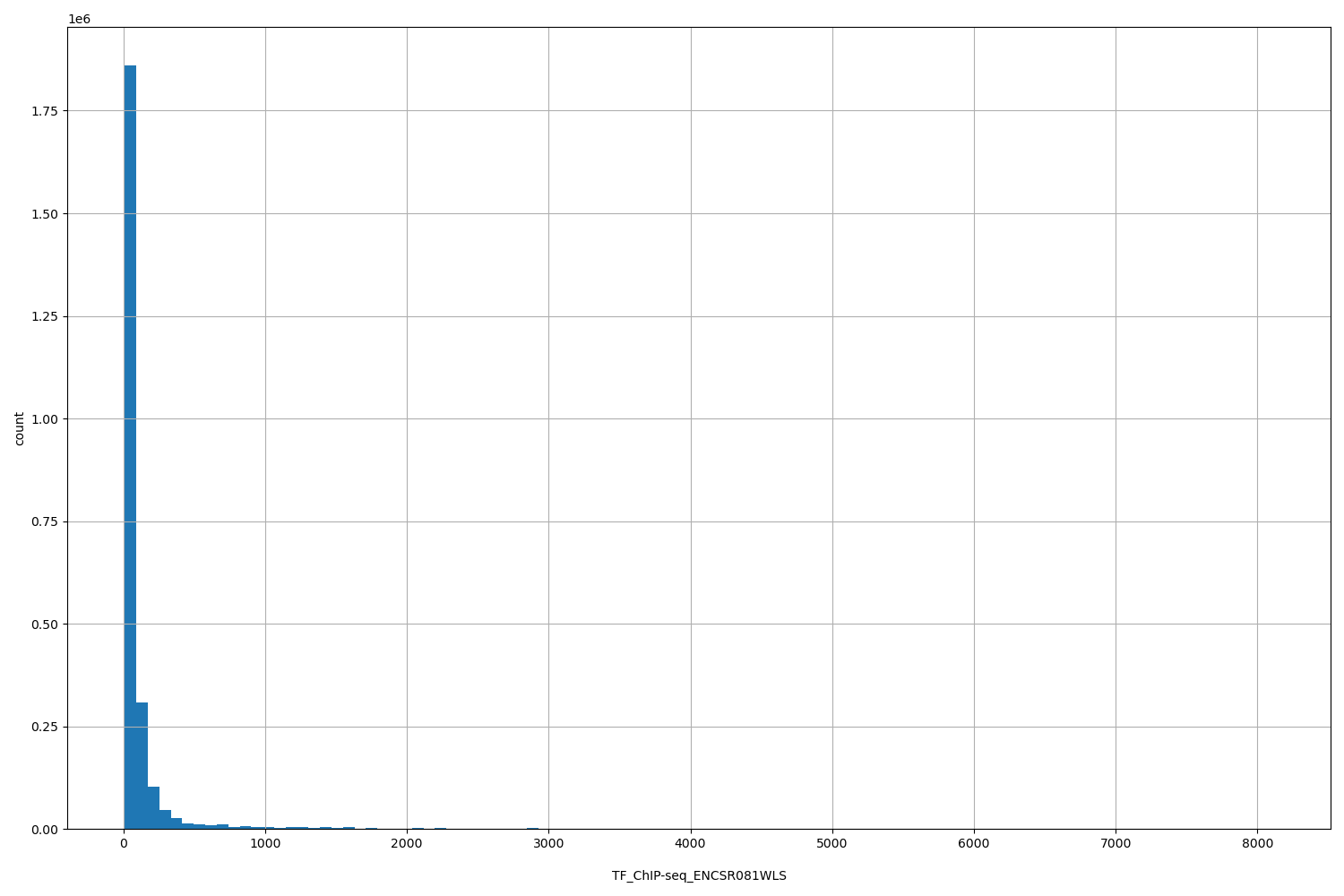
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR081WLS | float |
TF_ChIP-seq_ENCSR081WLS |
TF_ChIP-seq ENCSR081WLS [biosample_summary="Homo sapiens HepG2" and target="RAD51"]
|
 |
[11.4, 8.11e+03] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF681NJT.bed.gz | 138.63 KB | 02243fd2b0803e92a3cdaaa90f1f9c18 |
| ENCFF681NJT.bed.gz.dvc | 100.0 B | 8eb683a02878d6d578a1c25d345e7cb8 |
| ENCFF681NJT.tabix.bed.gz | 94.85 KB | 285bd2e6247ee990db74064bb6a144da |
| ENCFF681NJT.tabix.bed.gz.dvc | 105.0 B | 0d455532eea06f1d62846c8b63b78a1f |
| ENCFF681NJT.tabix.bed.gz.tbi | 72.06 KB | 38ef26f37fd0b72cbc6263118002f798 |
| ENCFF681NJT.tabix.bed.gz.tbi.dvc | 109.0 B | c6ec93d60eb08a7d14df1918a9f10012 |
| genomic_resource.yaml | 2.47 KB | 32985bcf5d9fe714e9781831fa22ea92 |
| genomic_resource_original.yaml | 2.38 KB | 698d13e948feb3b58e46d6828c87ac12 |
| statistics/ |