TF_ChIP-seq_ENCSR076SHT
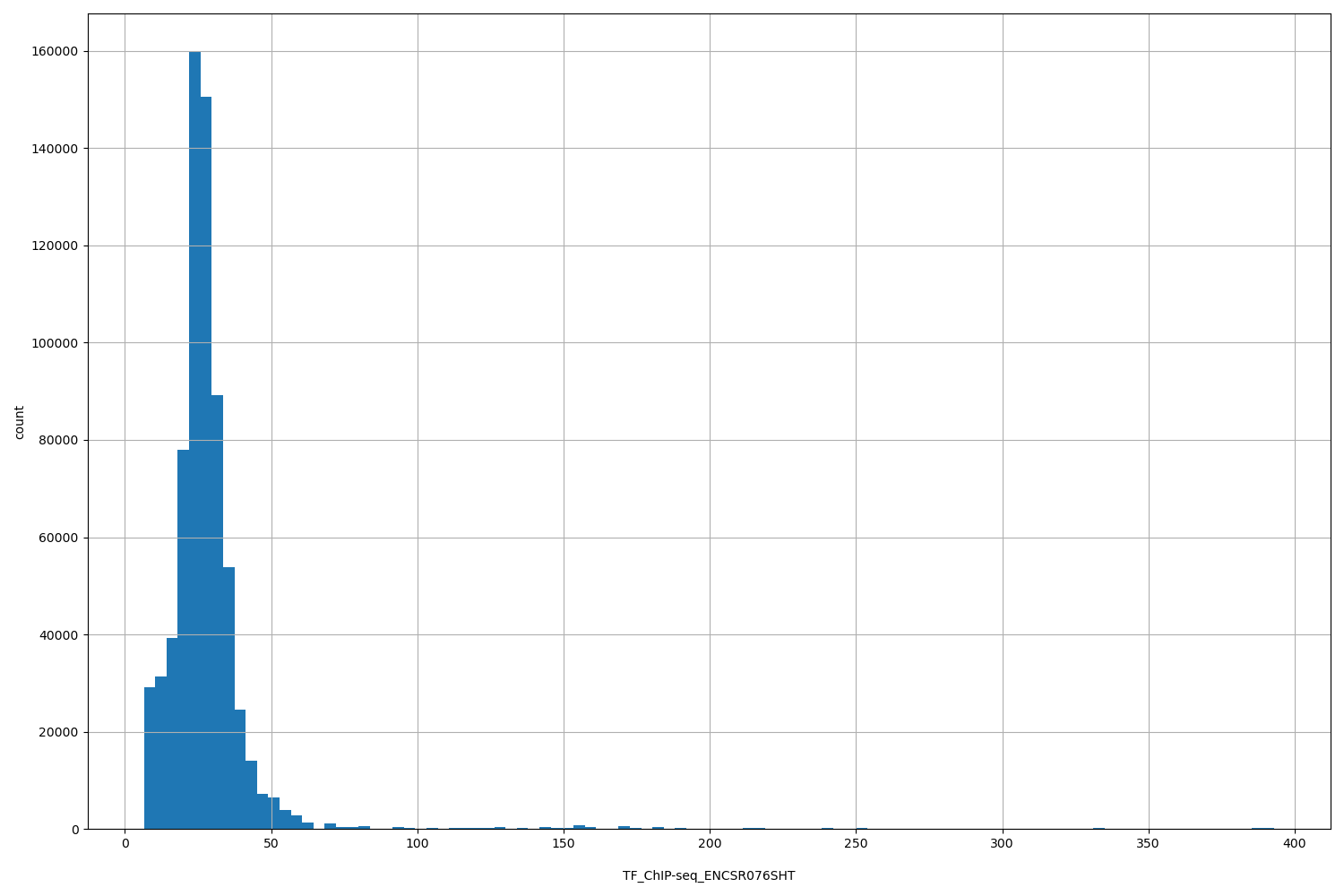
| Id: | TF_ChIP-seq/ENCSR076SHT |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR076SHT [biosamplesummary="Homo sapiens HepG2 genetically modified (insertion) using CRISPR targeting H. sapiens GPN1" and target="GPN1"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: genetically modified (insertion) using CRISPR targeting H. sapiens GPN1 output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN578KRU|/analyses/ENCAN578KRU/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_internal_action: Released analysis {ENCAN578KRU|/analyses/ENCAN578KRU/} has in progress subobject document {b32d4c51-de0d-483f-bf49-6ac9809ecd1d|/documents/b32d4c51-de0d-483f-bf49-6ac9809ecd1d/} audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF955LND|/files/ENCFF955LND/} processed by ChIP-seq ENCODE4 v1.5.1 GRCh38 pipeline have a rescue ratio of 1.34 and a self consistency ratio of 2.86. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF533NSU|/files/ENCFF533NSU/} processed by ChIP-seq ENCODE4 v1.5.1 GRCh38 pipeline have a rescue ratio of 1.34 and a self consistency ratio of 2.86. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR076SHT | float |
TF_ChIP-seq_ENCSR076SHT |
TF_ChIP-seq ENCSR076SHT [biosample_summary="Homo sapiens HepG2 genetically modified (insertion) using CRISPR targeting H. sapiens GPN1" and target="GPN1"]
|
 |
[6.51, 393] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF955LND.bed.gz | 44.1 KB | 075aec0bf5e24073d4342a873eea28c7 |
| ENCFF955LND.bed.gz.dvc | 99.0 B | 5fd0586239229cc4dbe3f40f47eac733 |
| ENCFF955LND.tabix.bed.gz | 30.95 KB | 71d2dea50a657cd3a842077ac91583b0 |
| ENCFF955LND.tabix.bed.gz.dvc | 105.0 B | c5613a83ebd36a591f64aef1fe2c24d4 |
| ENCFF955LND.tabix.bed.gz.tbi | 25.37 KB | de986370d35036a78f81b8b1fbc419ac |
| ENCFF955LND.tabix.bed.gz.tbi.dvc | 109.0 B | 7956490bee5e65525cbabae8f728467d |
| genomic_resource.yaml | 2.94 KB | 573d8300c4c12d8f6555a8f9b568c8ca |
| genomic_resource_original.yaml | 2.78 KB | c8af3d511444b6746c94b983b4aac6dd |
| statistics/ |