TF_ChIP-seq_ENCSR072GJV
| Id: | TF_ChIP-seq/ENCSR072GJV |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR072GJV [biosamplesummary="Homo sapiens HepG2 genetically modified (insertion) using CRISPR targeting H. sapiens HMG20A" and target="HMG20A"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: genetically modified (insertion) using CRISPR targeting H. sapiens HMG20A output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN373BEU|/analyses/ENCAN373BEU/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF298ZJA|/files/ENCFF298ZJA/}, {ENCFF187TBY|/files/ENCFF187TBY/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 1.90 and a self consistency ratio of 4.59. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF210MYZ|/files/ENCFF210MYZ/}, {ENCFF543WLU|/files/ENCFF543WLU/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 1.90 and a self consistency ratio of 4.59. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
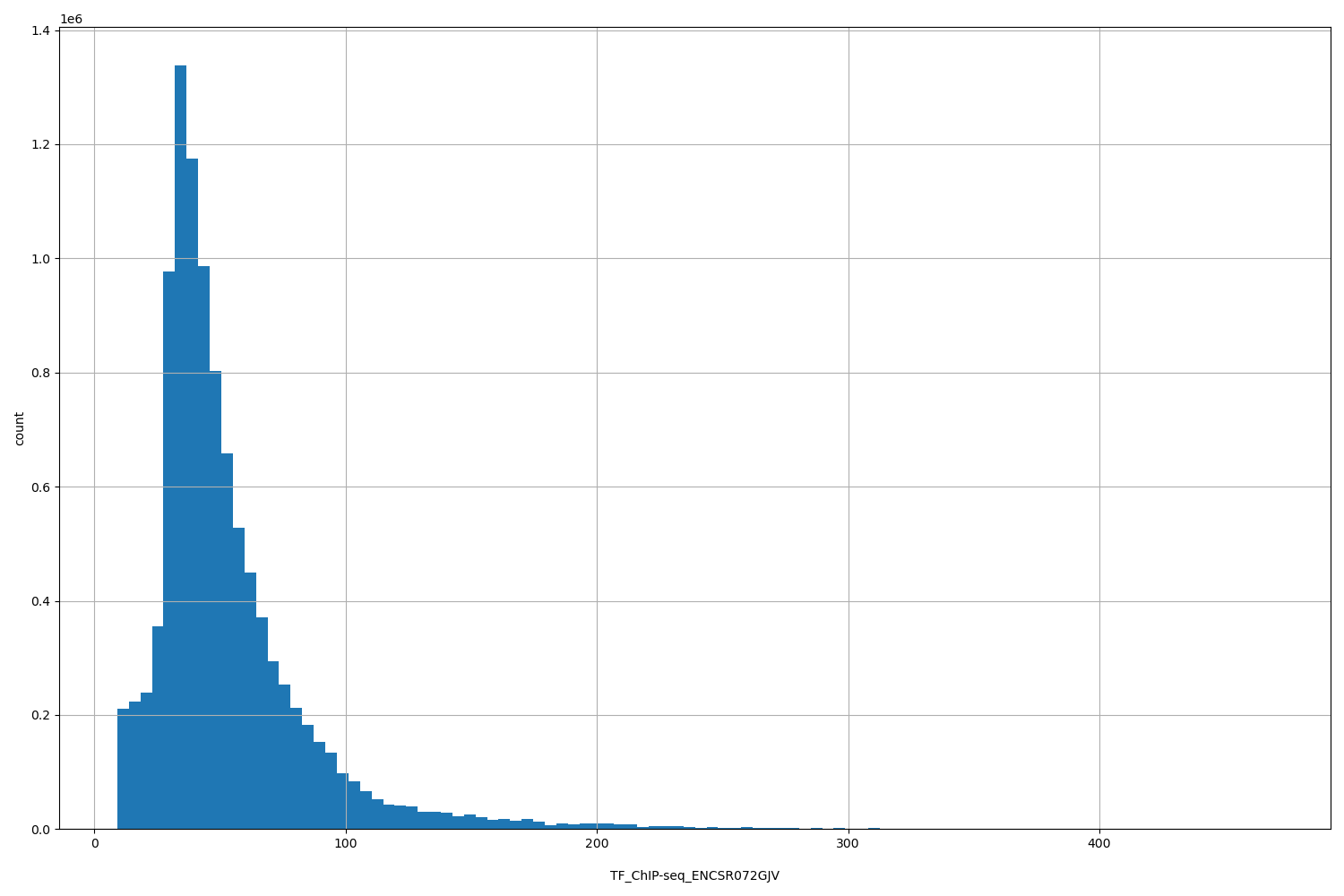
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR072GJV | float |
TF_ChIP-seq_ENCSR072GJV |
TF_ChIP-seq ENCSR072GJV [biosample_summary="Homo sapiens HepG2 genetically modified (insertion) using CRISPR targeting H. sapiens HMG20A" and target="HMG20A"]
|
 |
[9.14, 469] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF298ZJA.bed.gz | 436.68 KB | 0bdf7c5b32ecc67da08cc07185db3471 |
| ENCFF298ZJA.bed.gz.dvc | 100.0 B | b2f5b18153075c2876bcf258b2437620 |
| ENCFF298ZJA.tabix.bed.gz | 337.13 KB | c6119678dec3b011c25c146df0cf068a |
| ENCFF298ZJA.tabix.bed.gz.dvc | 106.0 B | 23e8b79cdd1d2ed8d7d044e01274e41e |
| ENCFF298ZJA.tabix.bed.gz.tbi | 150.65 KB | c91947533abb29f21eea2165245eba6d |
| ENCFF298ZJA.tabix.bed.gz.tbi.dvc | 110.0 B | 6906c18c6dd9a167588f3a60f4d94f80 |
| genomic_resource.yaml | 2.81 KB | 32ec4ca570130af844bba480ce44db62 |
| genomic_resource_original.yaml | 2.64 KB | 85fbf0e042d316c3790d119e4cc7ebe1 |
| statistics/ |