TF_ChIP-seq_ENCSR058DRG
| Id: | TF_ChIP-seq/ENCSR058DRG |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR058DRG [biosamplesummary="Homo sapiens K562 genetically modified (insertion) using CRISPR targeting H. sapiens CEBPG" and target="CEBPG"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: genetically modified (insertion) using CRISPR targeting H. sapiens CEBPG output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN159RRP|/analyses/ENCAN159RRP/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_warning: PBC1 (PCR Bottlenecking Coefficient 1, M1/M_distinct) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where some reads map (M_distinct). A PBC1 value in the range 0 - 0.5 is severe bottlenecking, 0.5 - 0.8 is moderate bottlenecking, 0.8 - 0.9 is mild bottlenecking, and > 0.9 is no bottlenecking. PBC1 value > 0.9 is recommended, but > 0.8 is acceptable. ENCODE processed alignments file {ENCFF611LFC|/files/ENCFF611LFC/} processed by ChIP-seq ENCODE4 v1.8.0 GRCh38 pipeline was generated from a library with PBC1 value of 0.90. audit_warning: PBC2 (PCR Bottlenecking Coefficient 2, M1/M2) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where two reads map uniquely (M2). A PBC2 value in the range 0 - 1 is severe bottlenecking, 1 - 3 is moderate bottlenecking, 3 - 10 is mild bottlenecking, > 10 is no bottlenecking. PBC2 value > 10 is recommended, but > 3 is acceptable. ENCODE processed alignments file {ENCFF611LFC|/files/ENCFF611LFC/} processed by ChIP-seq ENCODE4 v1.8.0 GRCh38 pipeline was generated from a library with PBC2 value of 9.63. audit_warning: PBC1 (PCR Bottlenecking Coefficient 1, M1/M_distinct) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where some reads map (M_distinct). A PBC1 value in the range 0 - 0.5 is severe bottlenecking, 0.5 - 0.8 is moderate bottlenecking, 0.8 - 0.9 is mild bottlenecking, and > 0.9 is no bottlenecking. PBC1 value > 0.9 is recommended, but > 0.8 is acceptable. ENCODE processed alignments file {ENCFF614LJY|/files/ENCFF614LJY/} processed by ChIP-seq ENCODE4 v1.8.0 GRCh38 pipeline was generated from a library with PBC1 value of 0.89. audit_warning: PBC2 (PCR Bottlenecking Coefficient 2, M1/M2) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where two reads map uniquely (M2). A PBC2 value in the range 0 - 1 is severe bottlenecking, 1 - 3 is moderate bottlenecking, 3 - 10 is mild bottlenecking, > 10 is no bottlenecking. PBC2 value > 10 is recommended, but > 3 is acceptable. ENCODE processed alignments file {ENCFF614LJY|/files/ENCFF614LJY/} processed by ChIP-seq ENCODE4 v1.8.0 GRCh38 pipeline was generated from a library with PBC2 value of 8.86. |
| Labels: |
|
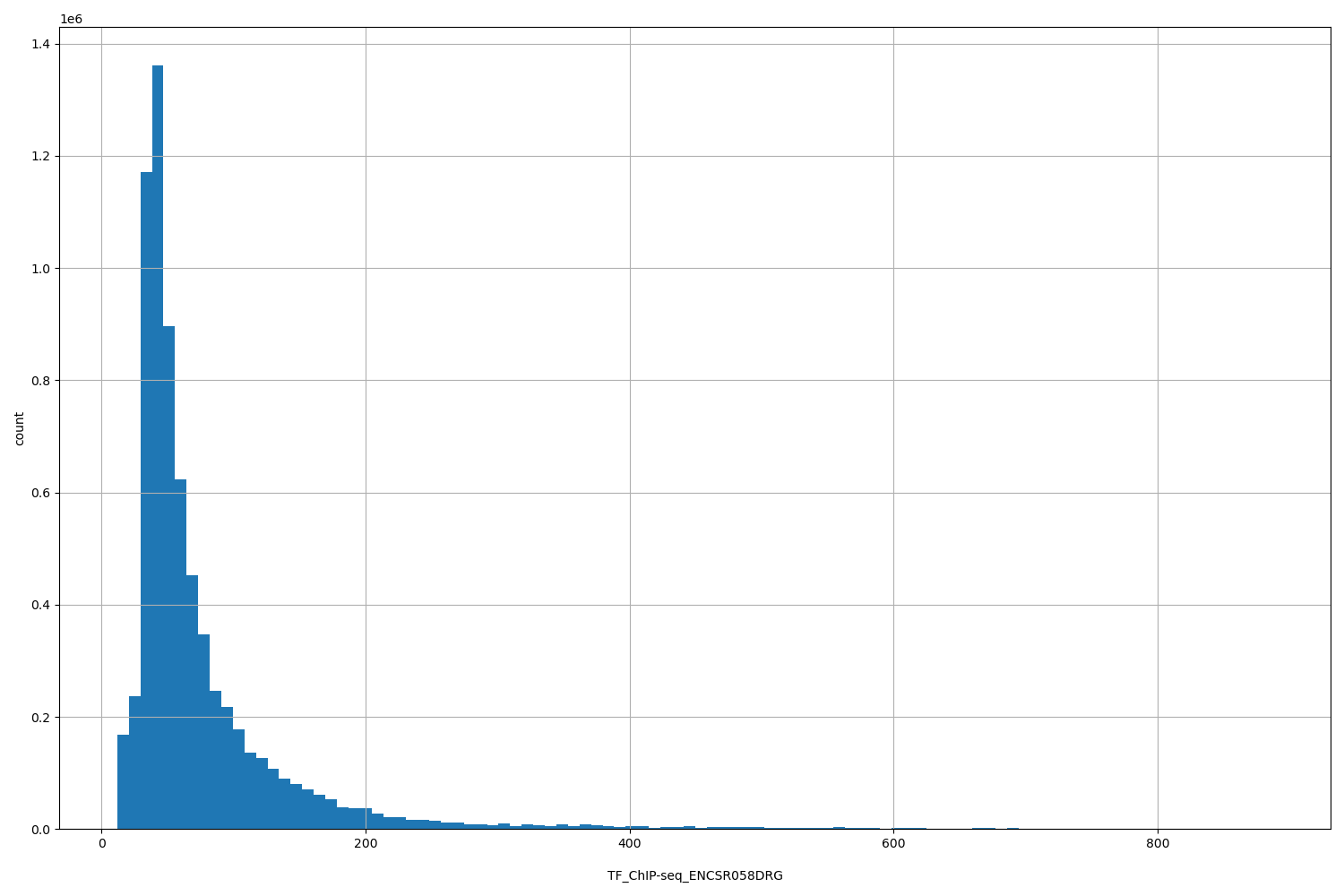
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR058DRG | float |
TF_ChIP-seq_ENCSR058DRG |
TF_ChIP-seq ENCSR058DRG [biosample_summary="Homo sapiens K562 genetically modified (insertion) using CRISPR targeting H. sapiens CEBPG" and target="CEBPG"]
|
 |
[12.1, 887] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF558AJI.bed.gz | 326.74 KB | 80d2e8c69d6dd36c8164a9c90a93367f |
| ENCFF558AJI.bed.gz.dvc | 100.0 B | 2eae896d893cf8c528ed4e6cd3f5af5e |
| ENCFF558AJI.tabix.bed.gz | 237.51 KB | ba044656977632f744d80b7f6874f5b7 |
| ENCFF558AJI.tabix.bed.gz.dvc | 106.0 B | b0a2109d06cfd8905774a57e257d9582 |
| ENCFF558AJI.tabix.bed.gz.tbi | 125.34 KB | 7239e379cfaf0150cdc8b6f79bf70f68 |
| ENCFF558AJI.tabix.bed.gz.tbi.dvc | 110.0 B | 5b8d11347bd4585f0b2b7f507b66b7b2 |
| genomic_resource.yaml | 4.23 KB | 5ac4393141f7102a795b11ea46aa2991 |
| genomic_resource_original.yaml | 4.07 KB | a630c7d7d063eba9038da36d28128307 |
| statistics/ |