TF_ChIP-seq_ENCSR048WVW
| Id: | TF_ChIP-seq/ENCSR048WVW |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR048WVW [biosamplesummary="Homo sapiens HepG2 genetically modified (insertion) using CRISPR targeting H. sapiens ZNF367" and target="ZNF367"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: genetically modified (insertion) using CRISPR targeting H. sapiens ZNF367 output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN294YTM|/analyses/ENCAN294YTM/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_internal_action: Released analysis {ENCAN294YTM|/analyses/ENCAN294YTM/} has in progress subobject document {c7255f44-3399-4535-b441-2ee372ad3be9|/documents/c7255f44-3399-4535-b441-2ee372ad3be9/} audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF236CPT|/files/ENCFF236CPT/} processed by ChIP-seq ENCODE4 v1.4.0 GRCh38 pipeline have a rescue ratio of 1.68 and a self consistency ratio of 4.34. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF673TZW|/files/ENCFF673TZW/} processed by ChIP-seq ENCODE4 v1.4.0 GRCh38 pipeline have a rescue ratio of 1.68 and a self consistency ratio of 4.34. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
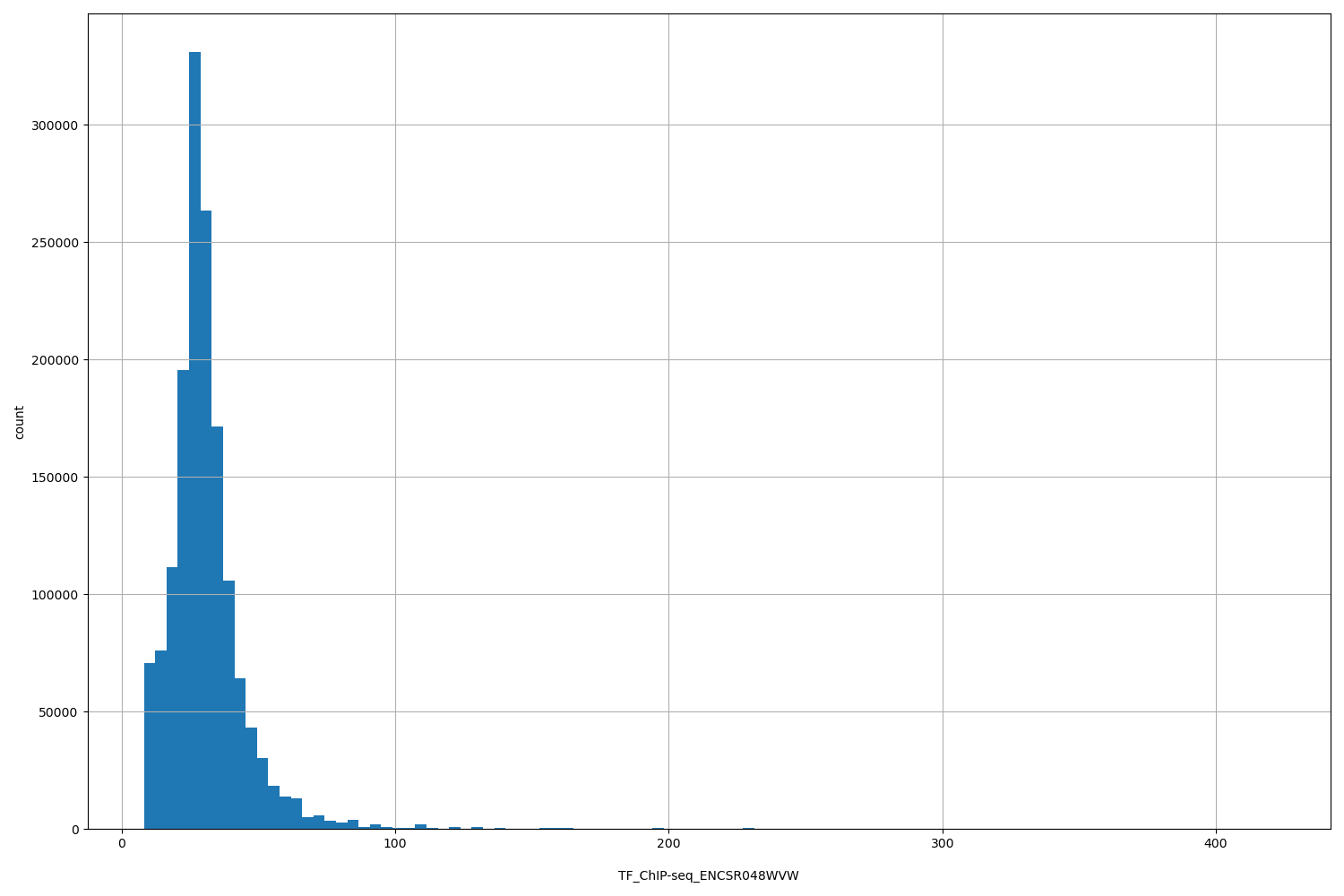
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR048WVW | float |
TF_ChIP-seq_ENCSR048WVW |
TF_ChIP-seq ENCSR048WVW [biosample_summary="Homo sapiens HepG2 genetically modified (insertion) using CRISPR targeting H. sapiens ZNF367" and target="ZNF367"]
|
 |
[8.14, 421] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF236CPT.bed.gz | 87.39 KB | 1659c386001bd29ef2805c96a03c2aae |
| ENCFF236CPT.bed.gz.dvc | 99.0 B | c761eb8348fcab8a7f4dbadee7ab894b |
| ENCFF236CPT.tabix.bed.gz | 60.69 KB | 5b6b714a64ecfce9717236a85a899dde |
| ENCFF236CPT.tabix.bed.gz.dvc | 105.0 B | f2d0e155517da39ffca280b77f6e846b |
| ENCFF236CPT.tabix.bed.gz.tbi | 40.06 KB | e1cef006ffab58a48ddbd1f80b1747f2 |
| ENCFF236CPT.tabix.bed.gz.tbi.dvc | 109.0 B | a8697a2b54d6660ecf40079caf289047 |
| genomic_resource.yaml | 2.96 KB | 408a52b3e6be46d3b771fcacb700893f |
| genomic_resource_original.yaml | 2.79 KB | b9206b112585a3143f4161b61bf073de |
| statistics/ |