TF_ChIP-seq_ENCSR036THH
| Id: | TF_ChIP-seq/ENCSR036THH |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR036THH [biosamplesummary="Homo sapiens SK-N-SH genetically modified\ \ (insertion) using CRISPR targeting H. sapiens BACH2 treated with 6 \u03BCM all-trans-retinoic\ \ acid for 48 hours" and target="BACH2"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: treated with 6 μM all-trans-retinoic acid for 48 hours genetically modified (insertion) using CRISPR targeting H. sapiens BACH2 output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN342HRW|/analyses/ENCAN342HRW/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_warning: PBC1 (PCR Bottlenecking Coefficient 1, M1/M_distinct) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where some reads map (M_distinct). A PBC1 value in the range 0 - 0.5 is severe bottlenecking, 0.5 - 0.8 is moderate bottlenecking, 0.8 - 0.9 is mild bottlenecking, and > 0.9 is no bottlenecking. PBC1 value > 0.9 is recommended, but > 0.8 is acceptable. ENCODE processed alignments file {ENCFF781EUO|/files/ENCFF781EUO/} processed by ChIP-seq ENCODE4 v1.9.0 GRCh38 pipeline was generated from a library with PBC1 value of 0.86. audit_warning: PBC2 (PCR Bottlenecking Coefficient 2, M1/M2) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where two reads map uniquely (M2). A PBC2 value in the range 0 - 1 is severe bottlenecking, 1 - 3 is moderate bottlenecking, 3 - 10 is mild bottlenecking, > 10 is no bottlenecking. PBC2 value > 10 is recommended, but > 3 is acceptable. ENCODE processed alignments file {ENCFF781EUO|/files/ENCFF781EUO/} processed by ChIP-seq ENCODE4 v1.9.0 GRCh38 pipeline was generated from a library with PBC2 value of 7.25. audit_warning: PBC1 (PCR Bottlenecking Coefficient 1, M1/M_distinct) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where some reads map (M_distinct). A PBC1 value in the range 0 - 0.5 is severe bottlenecking, 0.5 - 0.8 is moderate bottlenecking, 0.8 - 0.9 is mild bottlenecking, and > 0.9 is no bottlenecking. PBC1 value > 0.9 is recommended, but > 0.8 is acceptable. ENCODE processed alignments file {ENCFF955FIT|/files/ENCFF955FIT/} processed by ChIP-seq ENCODE4 v1.9.0 GRCh38 pipeline was generated from a library with PBC1 value of 0.86. audit_warning: PBC2 (PCR Bottlenecking Coefficient 2, M1/M2) is the ratio of the number of genomic locations where exactly one read maps uniquely (M1) to the number of genomic locations where two reads map uniquely (M2). A PBC2 value in the range 0 - 1 is severe bottlenecking, 1 - 3 is moderate bottlenecking, 3 - 10 is mild bottlenecking, > 10 is no bottlenecking. PBC2 value > 10 is recommended, but > 3 is acceptable. ENCODE processed alignments file {ENCFF955FIT|/files/ENCFF955FIT/} processed by ChIP-seq ENCODE4 v1.9.0 GRCh38 pipeline was generated from a library with PBC2 value of 7.20. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF947VJI|/files/ENCFF947VJI/} processed by ChIP-seq ENCODE4 v1.9.0 GRCh38 pipeline have a rescue ratio of 1.42 and a self consistency ratio of 3.10. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR036THH | float |
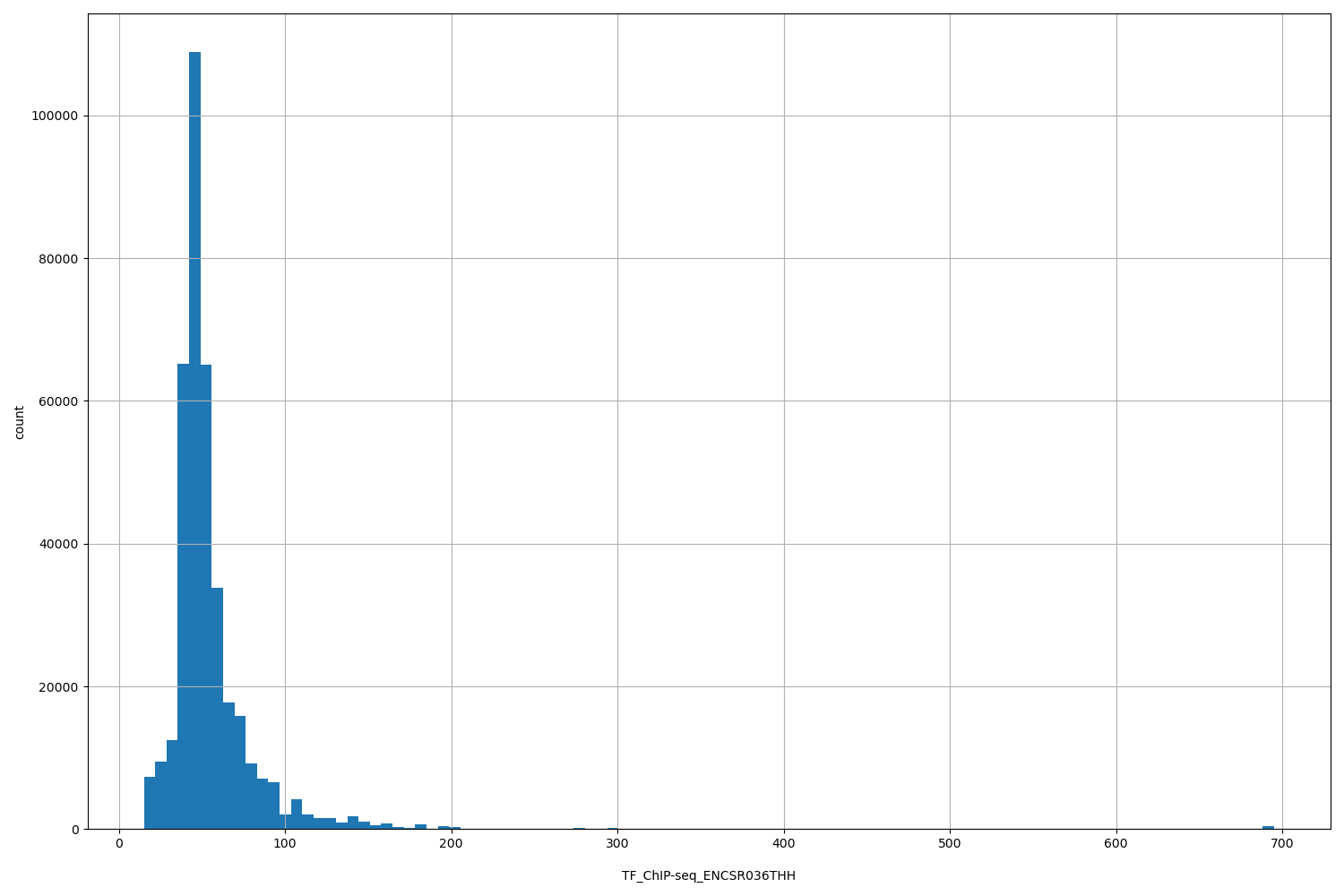
TF_ChIP-seq_ENCSR036THH |
TF_ChIP-seq ENCSR036THH [biosample_summary="Homo sapiens SK-N-SH genetically modified\ \ (insertion) using CRISPR targeting H. sapiens BACH2 treated with 6 \u03BCM all-trans-retinoic\ \ acid for 48 hours" and target="BACH2"]
|
 |
[15.1, 695] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF947VJI.bed.gz | 24.99 KB | f75f5083c221aa6a5cd2a745ed1abe9a |
| ENCFF947VJI.bed.gz.dvc | 99.0 B | e8c3786dfb3731ae0c325d1261b086a2 |
| ENCFF947VJI.tabix.bed.gz | 15.96 KB | f368c0a80f2d3eb09cbf07e58463a674 |
| ENCFF947VJI.tabix.bed.gz.dvc | 105.0 B | 338825e6a3727b1a02a351b551e971c3 |
| ENCFF947VJI.tabix.bed.gz.tbi | 17.32 KB | 9e0f9adb51499839c789a1503b0d1616 |
| ENCFF947VJI.tabix.bed.gz.tbi.dvc | 109.0 B | b2e1b7082c8784e3e4caf74811e3a68a |
| genomic_resource.yaml | 5.22 KB | 2789696c5e9fb25b7a73e2f3bc4c498b |
| genomic_resource_original.yaml | 4.99 KB | 6698dad75678c2de3e755efc21a76f7a |
| statistics/ |