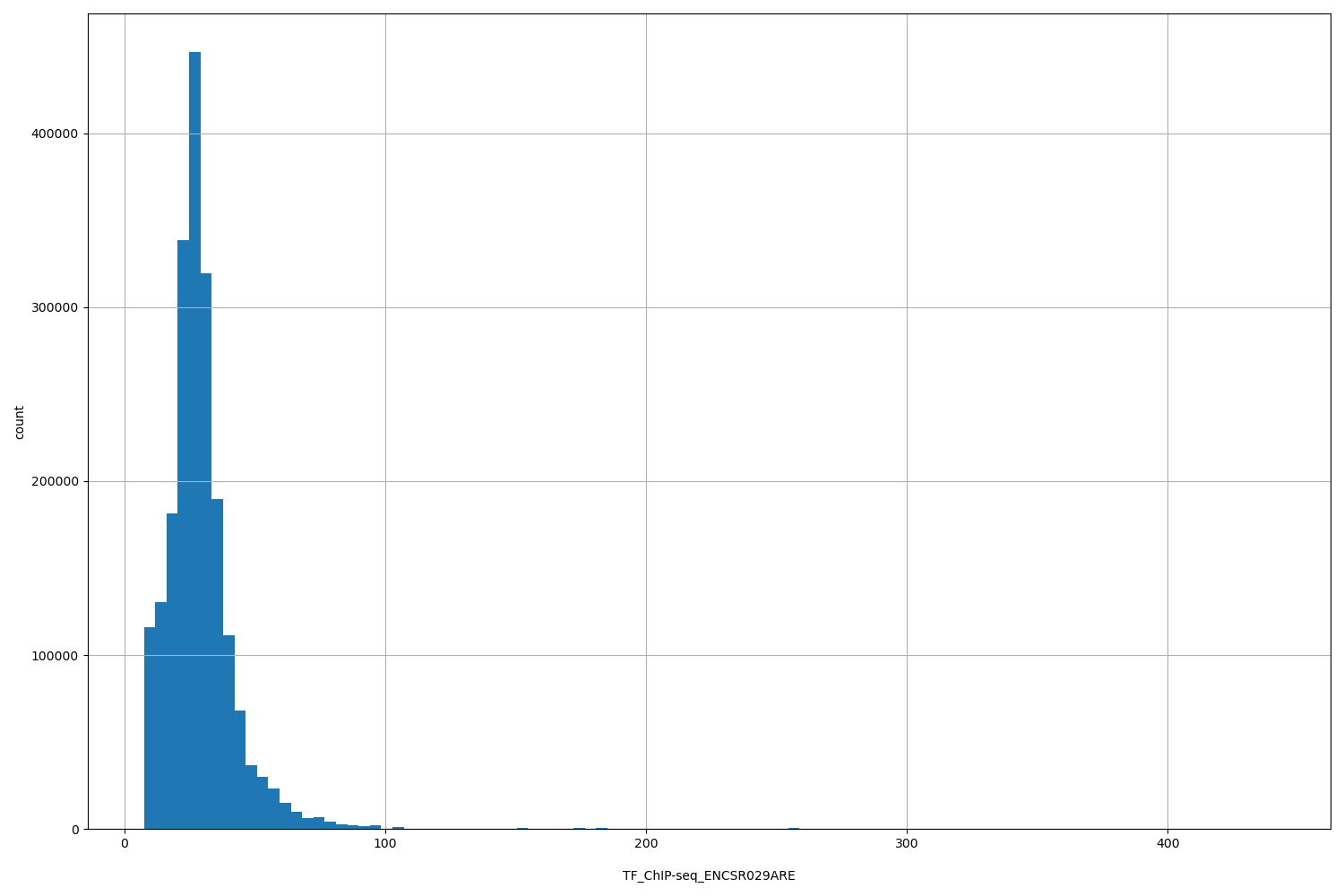
TF_ChIP-seq_ENCSR029ARE
| Id: | TF_ChIP-seq/ENCSR029ARE |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR029ARE [biosamplesummary="Homo sapiens HepG2 genetically modified (insertion) using CRISPR targeting H. sapiens ZNF773" and target="ZNF773"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: genetically modified (insertion) using CRISPR targeting H. sapiens ZNF773 output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN834UMJ|/analyses/ENCAN834UMJ/} has in progress subobject document {ff9c0567-71f2-45d2-bed4-5a2d34e380fc|/documents/ff9c0567-71f2-45d2-bed4-5a2d34e380fc/} audit_internal_action: Released analysis {ENCAN834UMJ|/analyses/ENCAN834UMJ/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF906WDG|/files/ENCFF906WDG/} processed by ChIP-seq ENCODE4 v1.4.0 GRCh38 pipeline have a rescue ratio of 1.67 and a self consistency ratio of 5.74. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF429EPY|/files/ENCFF429EPY/} processed by ChIP-seq ENCODE4 v1.4.0 GRCh38 pipeline have a rescue ratio of 1.67 and a self consistency ratio of 5.74. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR029ARE | float |
TF_ChIP-seq_ENCSR029ARE |
TF_ChIP-seq ENCSR029ARE [biosample_summary="Homo sapiens HepG2 genetically modified (insertion) using CRISPR targeting H. sapiens ZNF773" and target="ZNF773"]
|
 |
[7.58, 441] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF906WDG.bed.gz | 123.85 KB | eaa5d4a64cad7953cbda939cc275b558 |
| ENCFF906WDG.bed.gz.dvc | 100.0 B | 00fa5f943adc005b1ac1515ceb6c836f |
| ENCFF906WDG.tabix.bed.gz | 86.79 KB | 35266d52cf6f2972c43f0112cc1c60dd |
| ENCFF906WDG.tabix.bed.gz.dvc | 105.0 B | d2e98c8e5774870cc107bc93e1a73a83 |
| ENCFF906WDG.tabix.bed.gz.tbi | 52.96 KB | 9f5e284bc807826d983c0493d373cbc8 |
| ENCFF906WDG.tabix.bed.gz.tbi.dvc | 109.0 B | e7786ce541b170eff7391ebd9894c391 |
| genomic_resource.yaml | 2.96 KB | bfed14b541bc5ef55ce05aace9e64030 |
| genomic_resource_original.yaml | 2.79 KB | af4033c36c9b88d091476441cc4edcb3 |
| statistics/ |