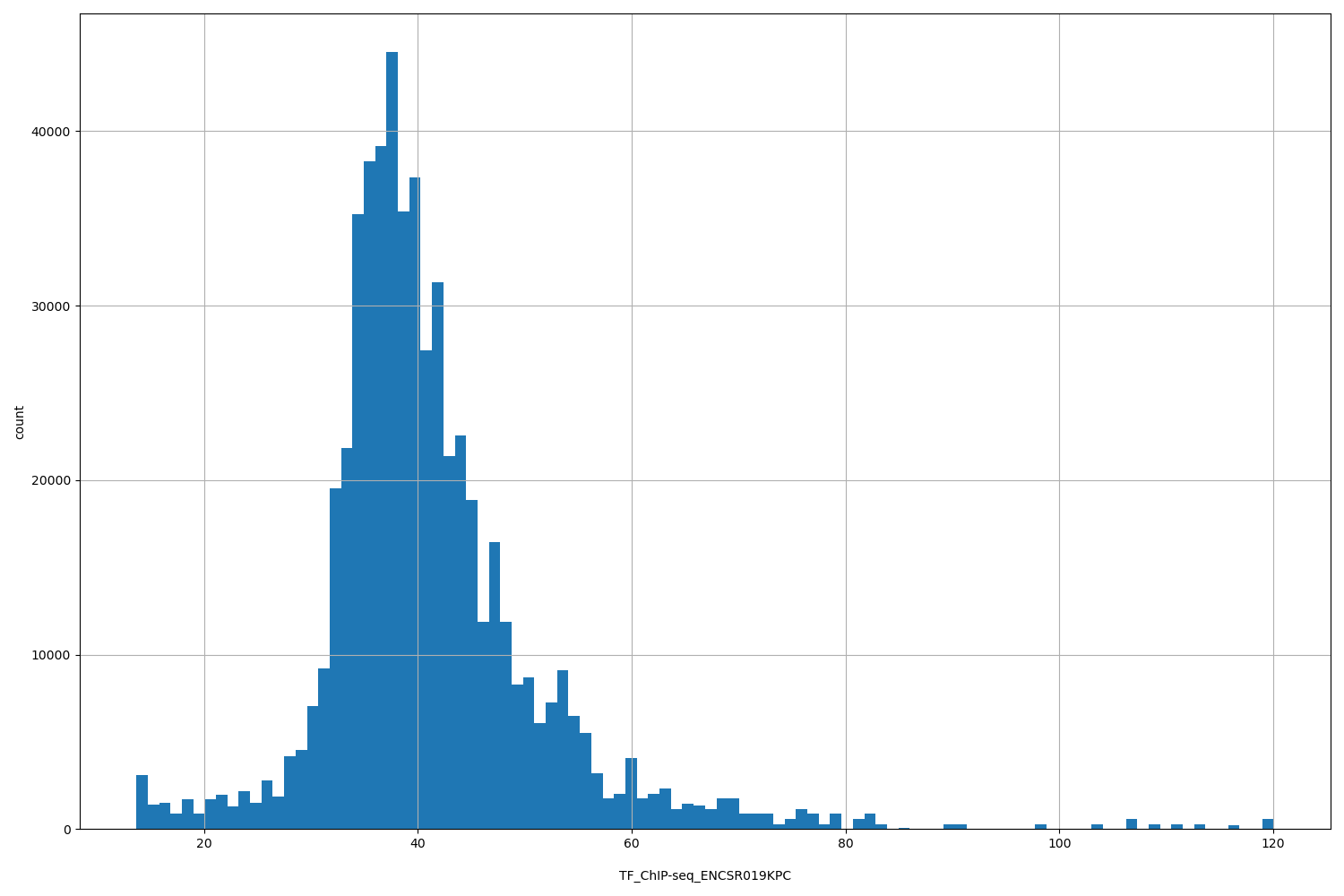
TF_ChIP-seq_ENCSR019KPC
| Id: | TF_ChIP-seq/ENCSR019KPC |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR019KPC [biosamplesummary="Homo sapiens HepG2" and target="PLRG1"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN891MYX|/analyses/ENCAN891MYX/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF926QHU|/files/ENCFF926QHU/}, {ENCFF276GUV|/files/ENCFF276GUV/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 2.75 and a self consistency ratio of 1.14. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF245EZL|/files/ENCFF245EZL/}, {ENCFF326RGO|/files/ENCFF326RGO/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 2.75 and a self consistency ratio of 1.14. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR019KPC | float |
TF_ChIP-seq_ENCSR019KPC |
TF_ChIP-seq ENCSR019KPC [biosample_summary="Homo sapiens HepG2" and target="PLRG1"]
|
 |
[13.7, 120] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF276GUV.bed.gz | 37.49 KB | e27ccc297cd456a07fa285e71f660549 |
| ENCFF276GUV.bed.gz.dvc | 99.0 B | 36a76812b130a638e38f7215acd0f5f3 |
| ENCFF276GUV.tabix.bed.gz | 23.92 KB | 3ed566beec852590f68943b930d85c22 |
| ENCFF276GUV.tabix.bed.gz.dvc | 105.0 B | 11b485f2d72a449c1707a8d788f65f1f |
| ENCFF276GUV.tabix.bed.gz.tbi | 22.23 KB | 62fb54df8a0cfd9536a5d411ab4bb3e8 |
| ENCFF276GUV.tabix.bed.gz.tbi.dvc | 109.0 B | 0e8c6589e9d8043944411cc146d172aa |
| genomic_resource.yaml | 2.47 KB | 75bee5e26199c92f11c77febbc959cc4 |
| genomic_resource_original.yaml | 2.38 KB | 9a02e93101628416eed6032e72292df8 |
| statistics/ |