TF_ChIP-seq_ENCSR011CIR
| Id: | TF_ChIP-seq/ENCSR011CIR |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR011CIR [biosamplesummary="Homo sapiens HepG2 genetically modified (insertion) using CRISPR targeting H. sapiens ZNF614" and target="ZNF614"] |
| Description: |
status: archived biological_replicates: Rep 1, Rep 2 summary: genetically modified (insertion) using CRISPR targeting H. sapiens ZNF614 output_type: optimal IDR thresholded peaks audit_internal_action: Archived analysis {ENCAN974IFN|/analyses/ENCAN974IFN/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF828NCK|/files/ENCFF828NCK/}, {ENCFF485ZUA|/files/ENCFF485ZUA/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 1.46 and a self consistency ratio of 2.42. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF733OUW|/files/ENCFF733OUW/}, {ENCFF944IVP|/files/ENCFF944IVP/} processed by ChIP-seq ENCODE3 hg19 pipeline have a rescue ratio of 1.46 and a self consistency ratio of 2.42. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
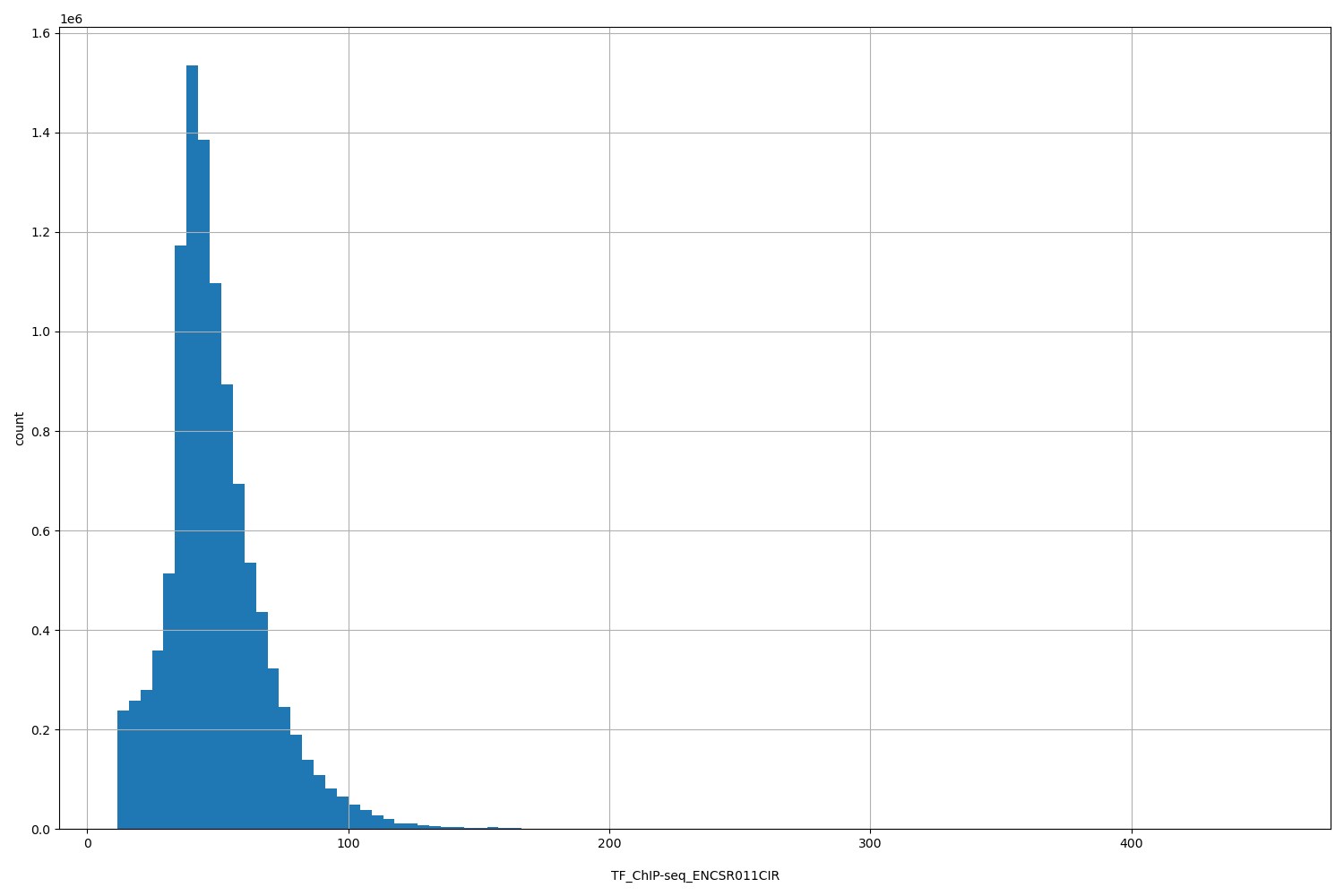
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR011CIR | float |
TF_ChIP-seq_ENCSR011CIR |
TF_ChIP-seq ENCSR011CIR [biosample_summary="Homo sapiens HepG2 genetically modified (insertion) using CRISPR targeting H. sapiens ZNF614" and target="ZNF614"]
|
 |
[11.5, 454] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF944IVP.bed.gz | 416.69 KB | 8f0cfb1c8ef37a3bb213a71519556f28 |
| ENCFF944IVP.bed.gz.dvc | 100.0 B | 0bd73304fee4dc87f0ee7b4c67878c4b |
| ENCFF944IVP.tabix.bed.gz | 319.4 KB | 46f4685f9176a79daa5c02214bee2956 |
| ENCFF944IVP.tabix.bed.gz.dvc | 106.0 B | ca5c08c3395bdd15d4a2888ad30b3a88 |
| ENCFF944IVP.tabix.bed.gz.tbi | 136.68 KB | 1b5f494f4bc00add733b09b63d1a1b26 |
| ENCFF944IVP.tabix.bed.gz.tbi.dvc | 110.0 B | bc4c9cf7ebc5b6f97662f9c464a4df6f |
| genomic_resource.yaml | 2.81 KB | 4e99d0b4d76094d4b664d4110f2cff91 |
| genomic_resource_original.yaml | 2.64 KB | f3cad9c9de950dbee98d1dd444912d99 |
| statistics/ |