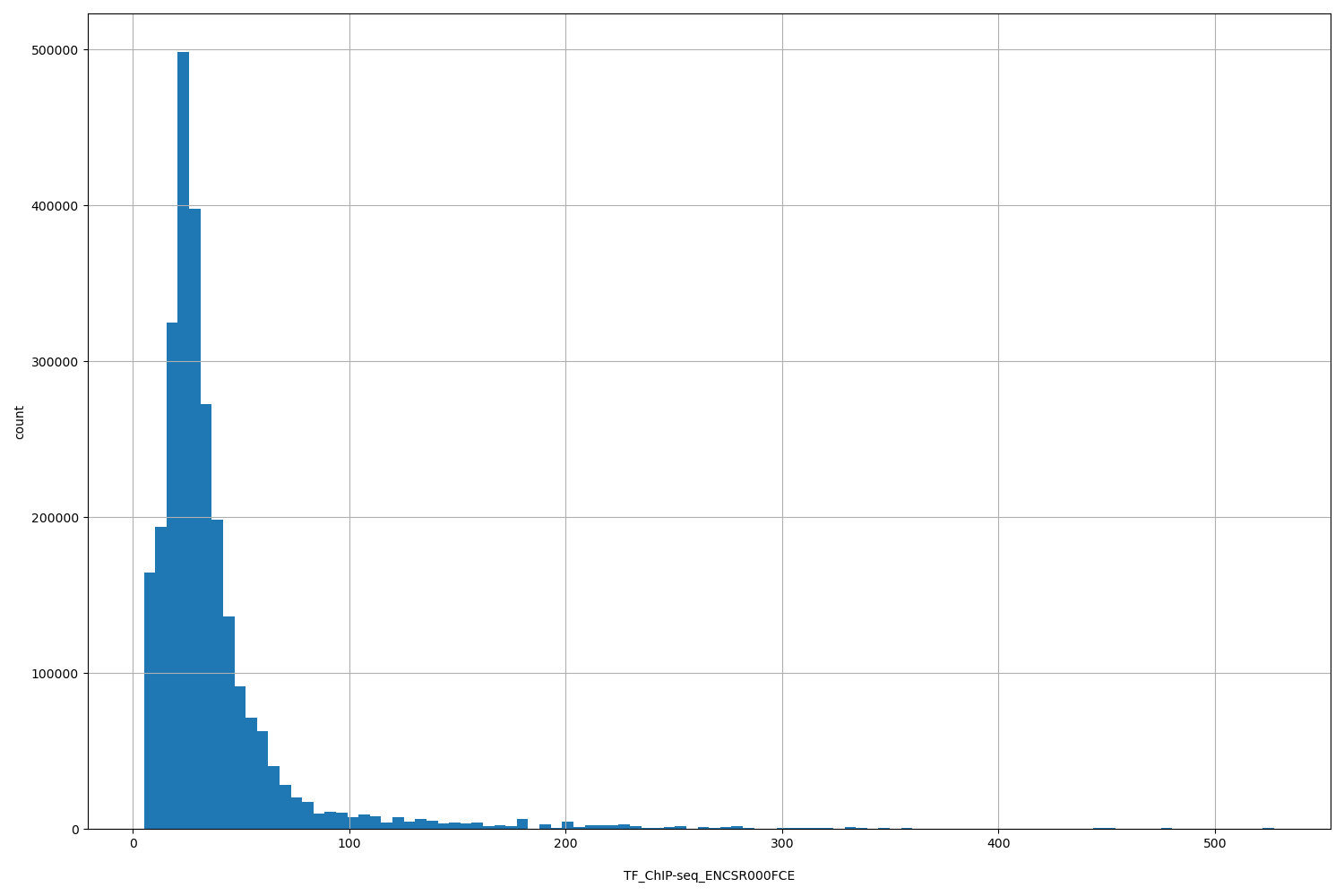
TF_ChIP-seq_ENCSR000FCE
| Id: | TF_ChIP-seq/ENCSR000FCE |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000FCE [biosamplesummary="Homo sapiens K562" and target="ETV6"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN420VBY|/analyses/ENCAN420VBY/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_internal_action: Released analysis {ENCAN420VBY|/analyses/ENCAN420VBY/} has in progress subobject document {746278b9-91d9-424a-9019-15242d7c3d71|/documents/746278b9-91d9-424a-9019-15242d7c3d71/} audit_warning: Processed alignments file {ENCFF779PUP|/files/ENCFF779PUP/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 18458202 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting ETV6-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Processed alignments file {ENCFF349ONJ|/files/ENCFF349ONJ/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 19437802 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting ETV6-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000FCE | float |
TF_ChIP-seq_ENCSR000FCE |
TF_ChIP-seq ENCSR000FCE [biosample_summary="Homo sapiens K562" and target="ETV6"]
|
 |
[5.09, 527] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF965PER.bed.gz | 120.3 KB | 0c6bff5143065945751d21983a178cfa |
| ENCFF965PER.bed.gz.dvc | 100.0 B | 1d3ac3d790786c6d5558fa068c337e0a |
| ENCFF965PER.tabix.bed.gz | 91.08 KB | b9278ab090174c6d1c1bca5c6416cb05 |
| ENCFF965PER.tabix.bed.gz.dvc | 105.0 B | 6119e7b8dcf9c103c43cf5696c39a791 |
| ENCFF965PER.tabix.bed.gz.tbi | 52.28 KB | 20cdf8179f073fed65152241c06af83d |
| ENCFF965PER.tabix.bed.gz.tbi.dvc | 109.0 B | 140813655de771b565a979cf0b48c211 |
| genomic_resource.yaml | 2.55 KB | b8e99af0463f5f32660326c0865f7b1b |
| genomic_resource_original.yaml | 2.46 KB | a1baa4b0daacc73707c2e9aeff0ec5c5 |
| statistics/ |