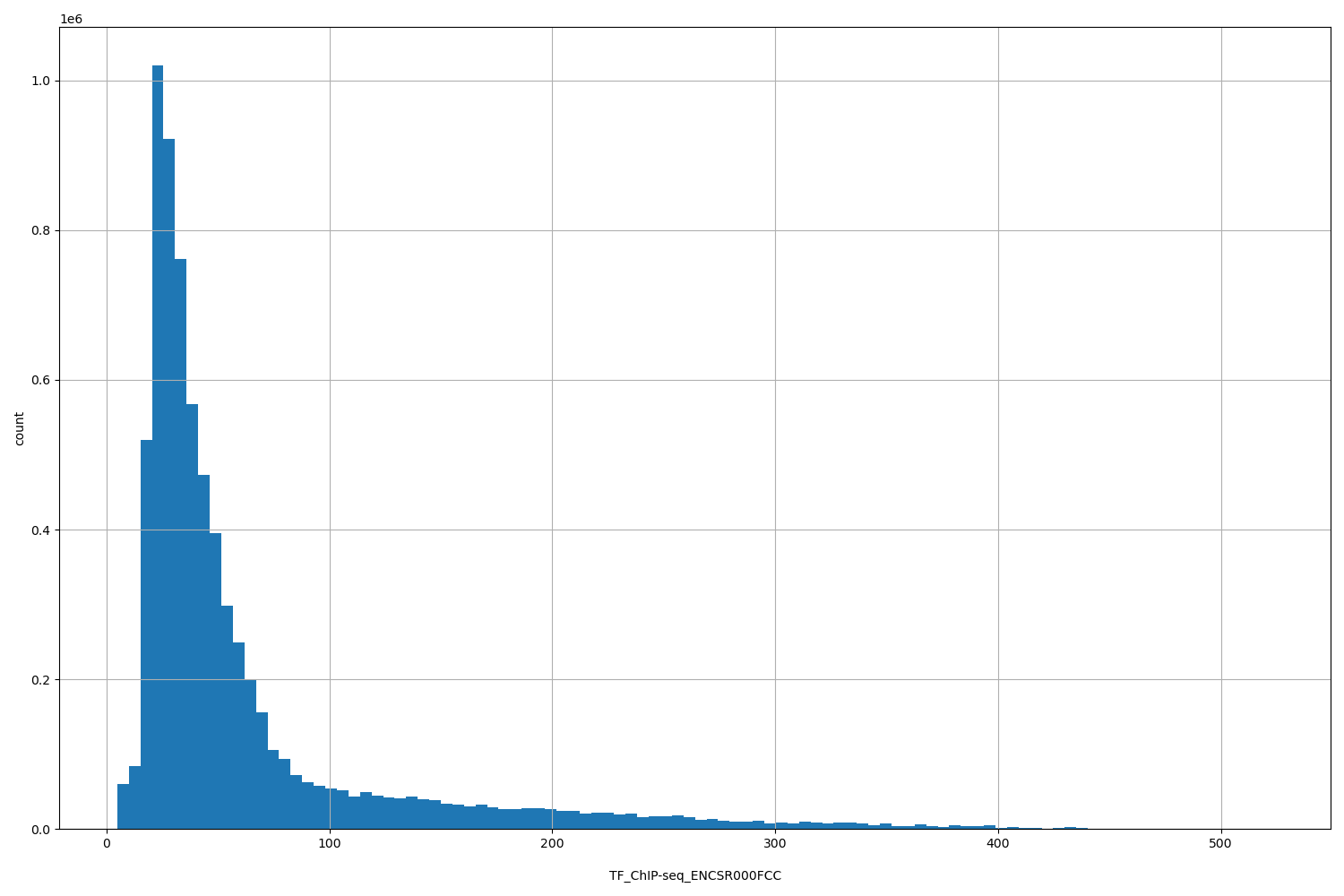
TF_ChIP-seq_ENCSR000FCC
| Id: | TF_ChIP-seq/ENCSR000FCC |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000FCC [biosamplesummary="Homo sapiens K562" and target="NFE2"] |
| Description: |
status: archived biological_replicates: Rep 1, Rep 2 summary: output_type: optimal IDR thresholded peaks audit_internal_action: Archived file {ENCFF312XHI|/files/ENCFF312XHI/} has replaced subobject file {5325a1e5-aca7-484b-a992-479fed28fbfa|/files/5325a1e5-aca7-484b-a992-479fed28fbfa/} audit_internal_action: Archived analysis {ENCAN245LAU|/analyses/ENCAN245LAU/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_warning: Processed alignments file {ENCFF198RTF|/files/ENCFF198RTF/} processed by ChIP-seq ENCODE3 GRCh38 pipeline has 18703425 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting NFE2-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000FCC | float |
TF_ChIP-seq_ENCSR000FCC |
TF_ChIP-seq ENCSR000FCC [biosample_summary="Homo sapiens K562" and target="NFE2"]
|
 |
[4.87, 523] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF312XHI.bed.gz | 484.77 KB | e9f3b1cbd2dddf9944661d8825db571a |
| ENCFF312XHI.bed.gz.dvc | 100.0 B | df9d6f71813beb8aed9b80c97ff54f85 |
| ENCFF312XHI.tabix.bed.gz | 362.03 KB | da393e45bed7ba705d02aa42a794a287 |
| ENCFF312XHI.tabix.bed.gz.dvc | 106.0 B | 5a160e0d9d59d469f818ac3f8276ca42 |
| ENCFF312XHI.tabix.bed.gz.tbi | 198.47 KB | d2872b5cd99df45f648c35ca3dc4fbeb |
| ENCFF312XHI.tabix.bed.gz.tbi.dvc | 110.0 B | b58aecb1dbabed8f2eb2e0c41b64da45 |
| genomic_resource.yaml | 2.04 KB | 8263e5457b16f4451971063c32e864c6 |
| genomic_resource_original.yaml | 1.94 KB | a60f268a46415d304ccec371ce59a221 |
| statistics/ |