TF_ChIP-seq_ENCSR000FAJ
| Id: | TF_ChIP-seq/ENCSR000FAJ |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000FAJ [biosamplesummary="Homo sapiens K562" and target="POLR2A"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN406CWX|/analyses/ENCAN406CWX/} has in progress subobject document {844cd949-6f5b-4bad-9d49-5d46f2b7733f|/documents/844cd949-6f5b-4bad-9d49-5d46f2b7733f/} audit_internal_action: Released analysis {ENCAN406CWX|/analyses/ENCAN406CWX/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_not_compliant: Processed alignments file {ENCFF380XSC|/files/ENCFF380XSC/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 6244819 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting POLR2A-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_not_compliant: Processed alignments file {ENCFF717URX|/files/ENCFF717URX/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 5864217 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting POLR2A-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
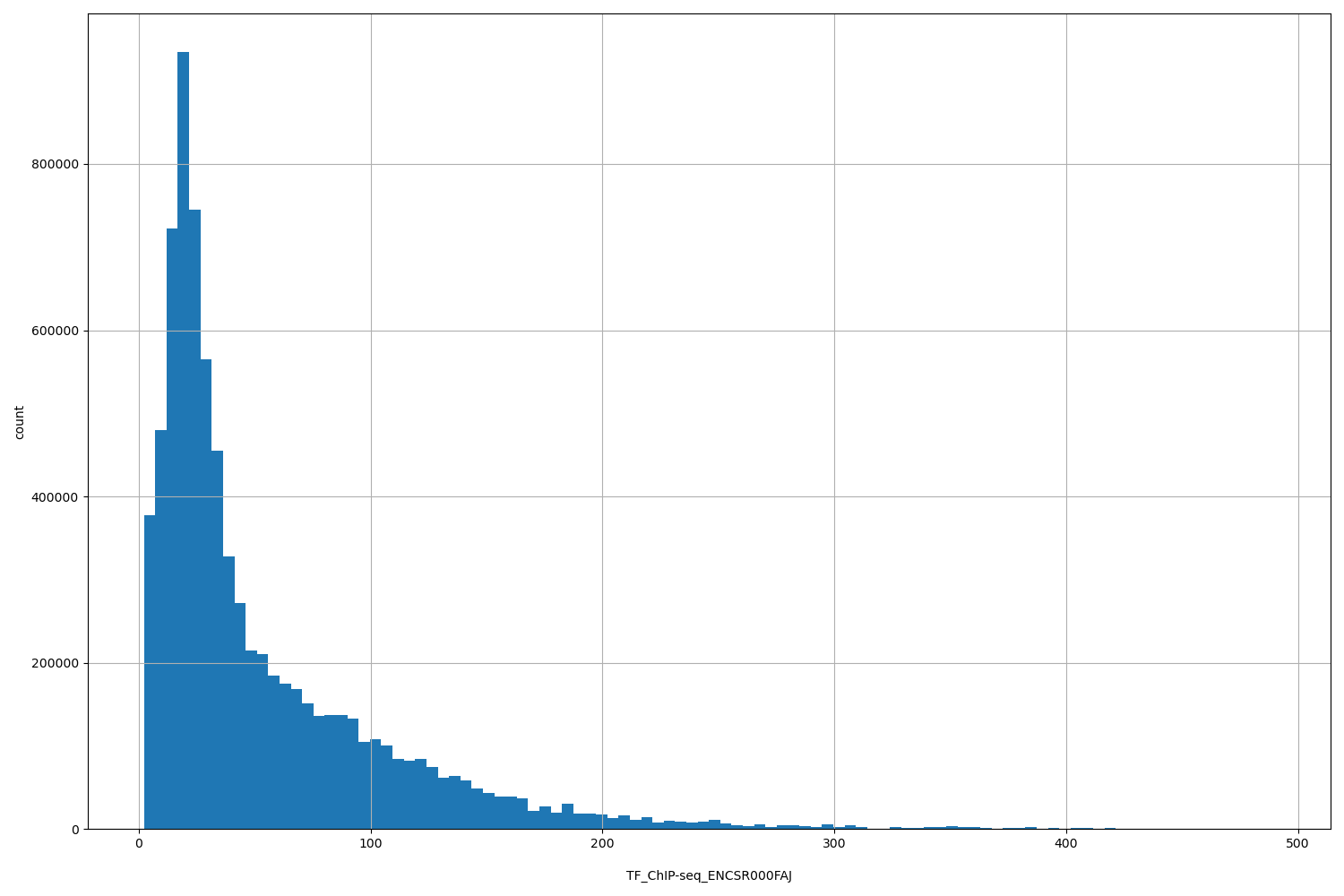
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000FAJ | float |
TF_ChIP-seq_ENCSR000FAJ |
TF_ChIP-seq ENCSR000FAJ [biosample_summary="Homo sapiens K562" and target="POLR2A"]
|
 |
[2.22, 490] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF107SJD.bed.gz | 389.1 KB | beb365ecb20e27529fcc392b762d81bf |
| ENCFF107SJD.bed.gz.dvc | 100.0 B | 070fe553e06199ac411d0b7e5e4d0b9d |
| ENCFF107SJD.tabix.bed.gz | 305.45 KB | 83f8b4786845b3c5acb96055b06bf338 |
| ENCFF107SJD.tabix.bed.gz.dvc | 106.0 B | 746c5d25e627499323322cbb51ec5312 |
| ENCFF107SJD.tabix.bed.gz.tbi | 116.89 KB | 22e939f65f6b376259521f4eccc264fe |
| ENCFF107SJD.tabix.bed.gz.tbi.dvc | 110.0 B | ccfb3a1b8a1b479b14cb0f582eb71246 |
| genomic_resource.yaml | 2.58 KB | 898a5a6746c6ad7edbed131b4ee547bc |
| genomic_resource_original.yaml | 2.48 KB | 2698a8b5656f7b61744c360f138aa1ce |
| statistics/ |