TF_ChIP-seq_ENCSR000FAE
TF_ChIP-seq ENCSR000FAE [biosample_summary="Homo sapiens K562" and target="MAX"]
| Id: | TF_ChIP-seq/ENCSR000FAE |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000FAE [biosamplesummary="Homo sapiens K562" and target="MAX"] |
| Description: |
status: released biological_replicates: summary: output_type: peaks audit_internal_action: derived_from is a list of files that were used to create a given file; for example, fastq file(s) will appear in the derived_from list of an alignments file. Processed file {ENCFF001VNT|/files/ENCFF001VNT/} is missing the requisite file specification in its derived_from list. |
| Labels: |
|
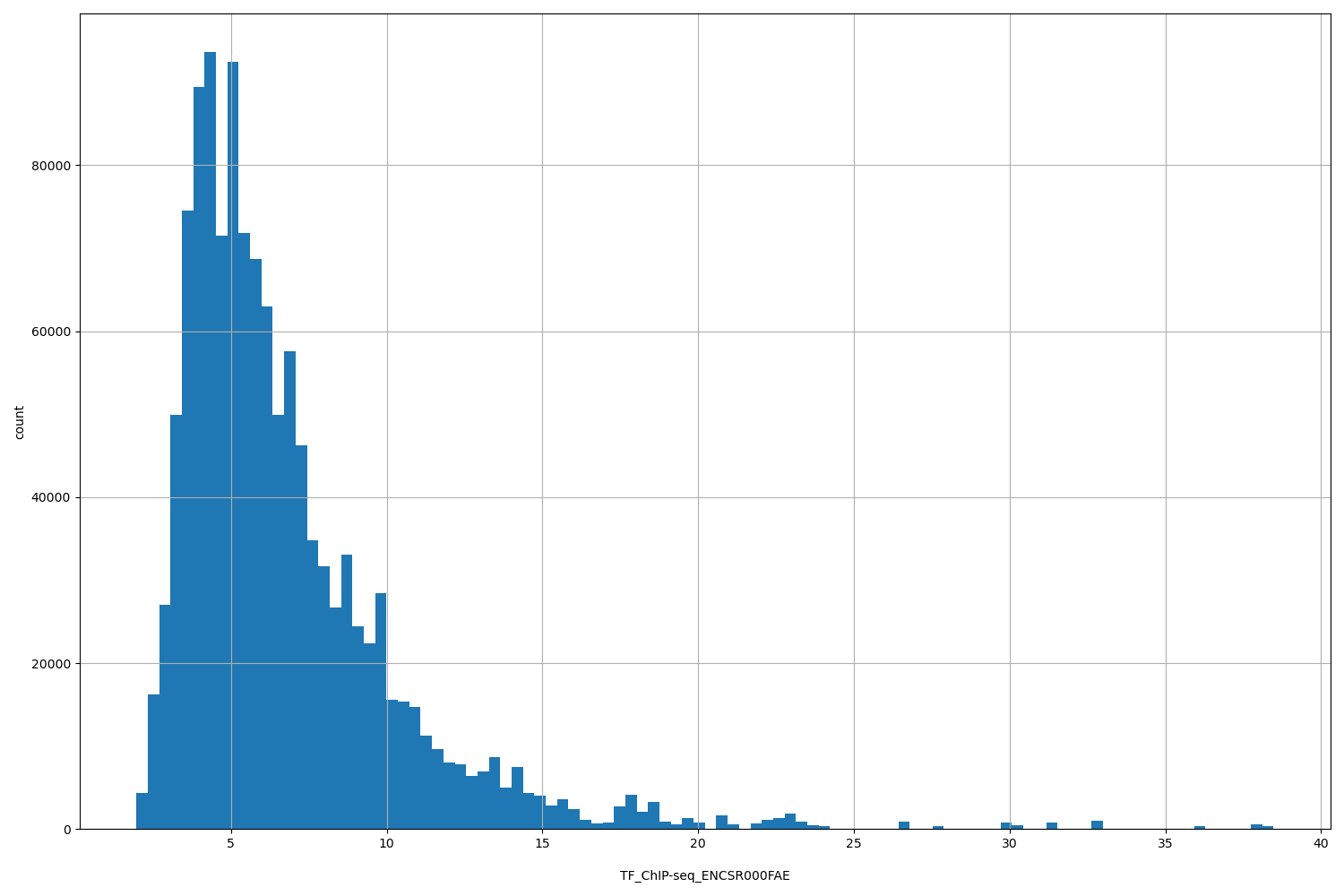
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000FAE | float |
TF_ChIP-seq_ENCSR000FAE |
TF_ChIP-seq ENCSR000FAE [biosample_summary="Homo sapiens K562" and target="MAX"]
|
 |
[1.96, 38.5] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF001VNT.bed.gz | 44.51 KB | a19aaf8abd95dbbb946feef40beb6e21 |
| ENCFF001VNT.bed.gz.dvc | 99.0 B | a8bb3d641007c6a18c634680fac37862 |
| ENCFF001VNT.tabix.bed.gz | 21.15 KB | 5703a2faa02e04e5c2a6891266e2ce57 |
| ENCFF001VNT.tabix.bed.gz.dvc | 105.0 B | aafae0008e232694f2624aed439927f2 |
| ENCFF001VNT.tabix.bed.gz.tbi | 23.7 KB | c7952fa973dba0ab6153caa50c4b59ff |
| ENCFF001VNT.tabix.bed.gz.tbi.dvc | 109.0 B | 4cb7faed71c352266c78dca75039fa5d |
| genomic_resource.yaml | 1.44 KB | 2e3fcdc8871ff98297324163595d2aaf |
| genomic_resource_original.yaml | 1.35 KB | c4275425f590fc9796e8238b909ef1ca |
| statistics/ |