TF_ChIP-seq_ENCSR000FAC
TF_ChIP-seq ENCSR000FAC [biosample_summary="Homo sapiens K562" and target="XRCC4"]
| Id: | TF_ChIP-seq/ENCSR000FAC |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000FAC [biosamplesummary="Homo sapiens K562" and target="XRCC4"] |
| Description: |
status: released biological_replicates: summary: output_type: peaks audit_internal_action: derived_from is a list of files that were used to create a given file; for example, fastq file(s) will appear in the derived_from list of an alignments file. Processed file {ENCFF001VPI|/files/ENCFF001VPI/} is missing the requisite file specification in its derived_from list. |
| Labels: |
|
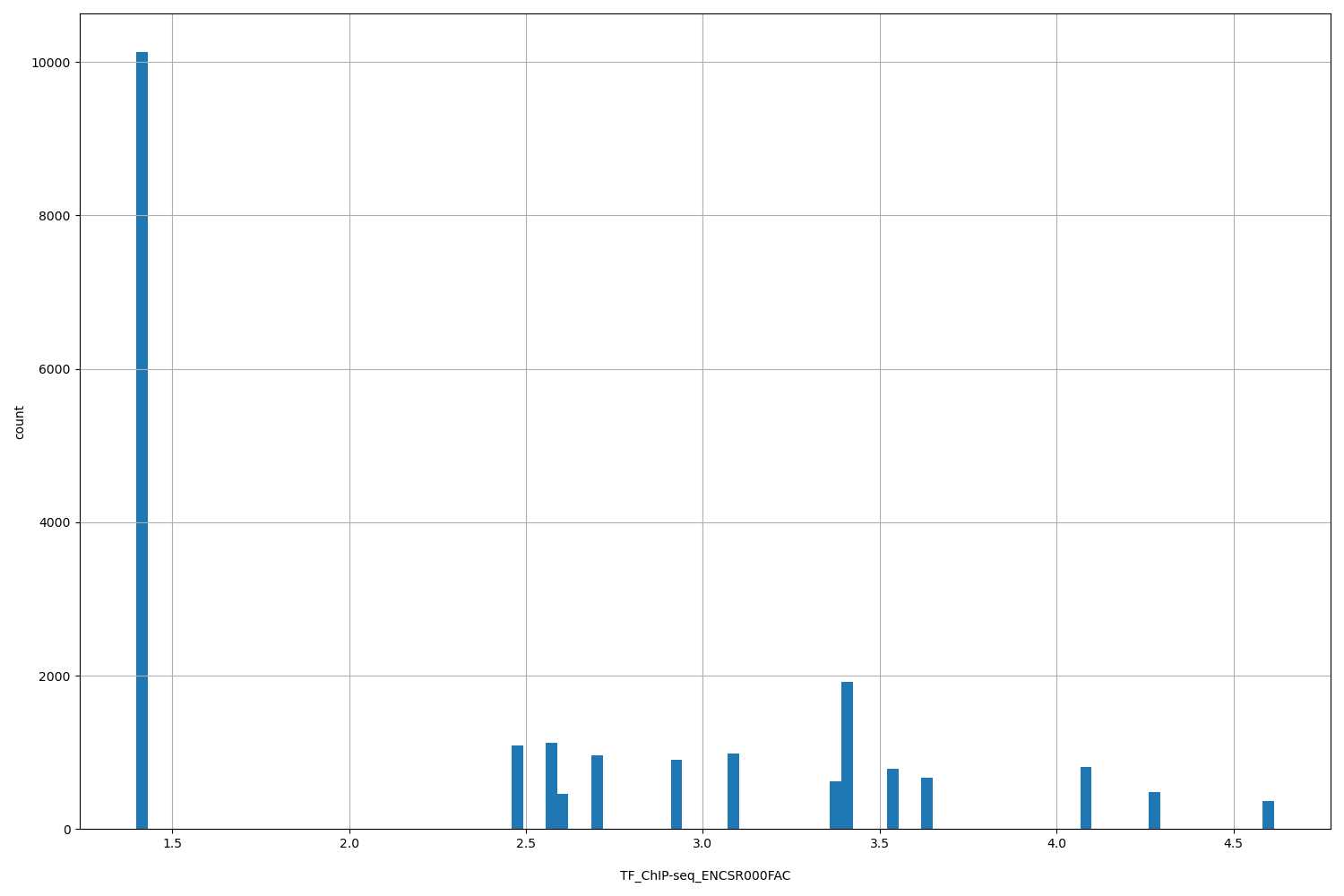
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000FAC | float |
TF_ChIP-seq_ENCSR000FAC |
TF_ChIP-seq ENCSR000FAC [biosample_summary="Homo sapiens K562" and target="XRCC4"]
|
 |
[1.4, 4.61] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF001VPI.bed.gz | 536.0 B | 53b582d7e33cebdad006cc70b1648caf |
| ENCFF001VPI.bed.gz.dvc | 97.0 B | fd4a61f3450086b2676293bffc1ad430 |
| ENCFF001VPI.tabix.bed.gz | 258.0 B | c056e5c0b49a3104d764791df84d5fa5 |
| ENCFF001VPI.tabix.bed.gz.dvc | 103.0 B | a562179118246521ade43eea085c24bd |
| ENCFF001VPI.tabix.bed.gz.tbi | 1.19 KB | 39241ee3b4bf2057ef4e44d1dbf2513e |
| ENCFF001VPI.tabix.bed.gz.tbi.dvc | 108.0 B | e7314c3e1b94cfad87ca22b1232a60ac |
| genomic_resource.yaml | 1.45 KB | bd588d72744f81bc89172327151d2b34 |
| genomic_resource_original.yaml | 1.35 KB | 1fabbd55c49c434648c0256f1627199f |
| statistics/ |