TF_ChIP-seq_ENCSR000EZF
| Id: | TF_ChIP-seq/ENCSR000EZF |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000EZF [biosamplesummary="Homo sapiens HeLa-S3" and target="MAX"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: conservative IDR thresholded peaks audit_error: Processed alignments file {ENCFF920GKG|/files/ENCFF920GKG/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 4600979 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting MAX-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_error: Processed alignments file {ENCFF072SJG|/files/ENCFF072SJG/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 4187535 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting MAX-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_internal_action: Released analysis {ENCAN510KGU|/analyses/ENCAN510KGU/} has in progress subobject document {8b033911-c3b0-4b51-b3d1-e9560c2eba54|/documents/8b033911-c3b0-4b51-b3d1-e9560c2eba54/} audit_internal_action: Released analysis {ENCAN510KGU|/analyses/ENCAN510KGU/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} |
| Labels: |
|
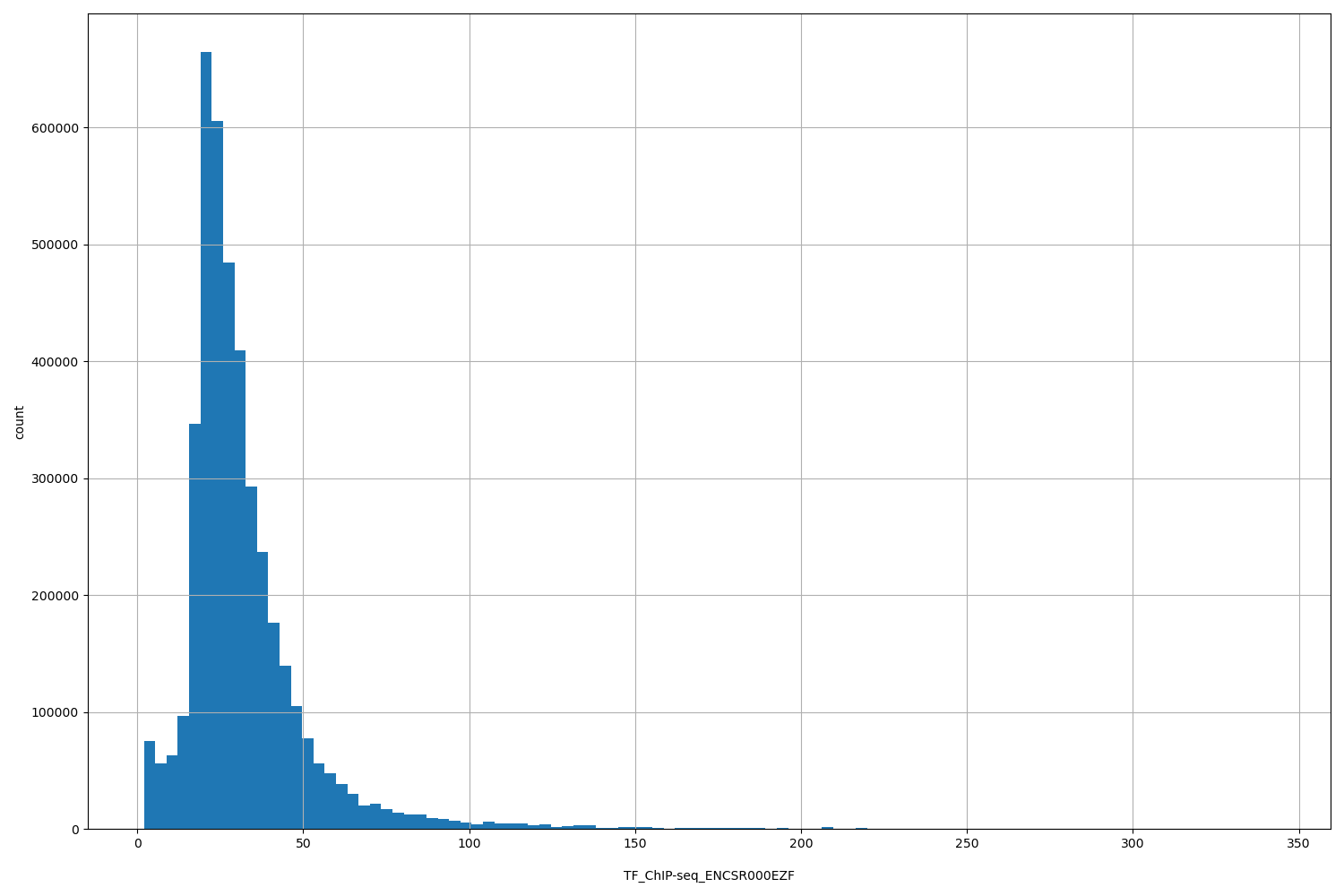
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000EZF | float |
TF_ChIP-seq_ENCSR000EZF |
TF_ChIP-seq ENCSR000EZF [biosample_summary="Homo sapiens HeLa-S3" and target="MAX"]
|
 |
[2, 343] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF123NLG.bed.gz | 220.85 KB | d8b4d9fd7d423a85cbf84b7f10ba33fe |
| ENCFF123NLG.bed.gz.dvc | 100.0 B | 3f4c074461208634634a5b605859b148 |
| ENCFF123NLG.tabix.bed.gz | 171.92 KB | 739a1ee3f4f86d246126a40131b6653e |
| ENCFF123NLG.tabix.bed.gz.dvc | 106.0 B | 04beb596fc3664e9dd7a92248b7dae97 |
| ENCFF123NLG.tabix.bed.gz.tbi | 99.27 KB | 95fca7d4ee5ecaed2e14284c2280858c |
| ENCFF123NLG.tabix.bed.gz.tbi.dvc | 110.0 B | 82b0d567b1d04b28d26e88457579b072 |
| genomic_resource.yaml | 2.57 KB | ee32ae988170d26d6782e54fb618c7bd |
| genomic_resource_original.yaml | 2.47 KB | 470e696d8bfbd850a6827d8568d2dd96 |
| statistics/ |