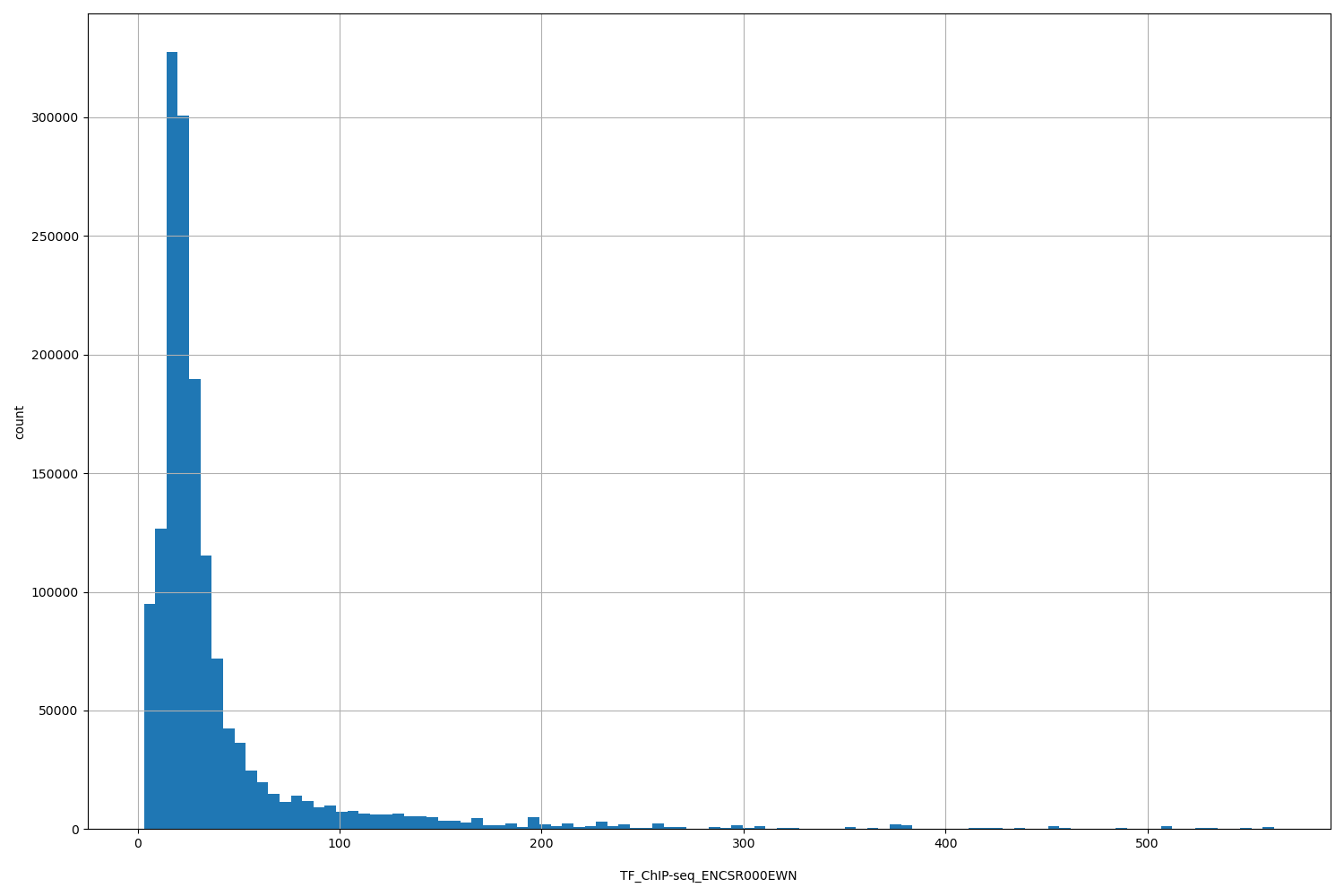
TF_ChIP-seq_ENCSR000EWN
| Id: | TF_ChIP-seq/ENCSR000EWN |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000EWN [biosamplesummary="Homo sapiens K562" and target="ZNF263"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: optimal IDR thresholded peaks audit_internal_action: File {ENCFF579ETM|/files/ENCFF579ETM/} with status 'released' is derived from file {ENCFF419RSJ|/files/ENCFF419RSJ/} with status 'archived'. audit_internal_action: Archived analysis {ENCAN506IUH|/analyses/ENCAN506IUH/} has in progress subobject quality standard {encode3-tf-chip|/quality-standards/encode3-tf-chip/} audit_error: Processed alignments file {ENCFF891GFG|/files/ENCFF891GFG/} processed by ChIP-seq ENCODE3 GRCh38 pipeline has 4555088 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting ZNF263-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_error: Processed alignments file {ENCFF094FZQ|/files/ENCFF094FZQ/} processed by ChIP-seq ENCODE3 GRCh38 pipeline has 4311179 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting ZNF263-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000EWN | float |
TF_ChIP-seq_ENCSR000EWN |
TF_ChIP-seq ENCSR000EWN [biosample_summary="Homo sapiens K562" and target="ZNF263"]
|
 |
[3.05, 563] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF579ETM.bed.gz | 66.2 KB | b6c38bae2ef61a659aadf55b977a891a |
| ENCFF579ETM.bed.gz.dvc | 99.0 B | 7706b73e13eba5c463044a38c5c39c80 |
| ENCFF579ETM.tabix.bed.gz | 53.67 KB | b1abc6cb64e57306bc61b1f70d600b30 |
| ENCFF579ETM.tabix.bed.gz.dvc | 105.0 B | 83748cf936314c904e1a416a886df3f8 |
| ENCFF579ETM.tabix.bed.gz.tbi | 34.6 KB | 20f745903ed80922f2c431a9c6b21148 |
| ENCFF579ETM.tabix.bed.gz.tbi.dvc | 109.0 B | 4dd950c01b918c3cbc35cf0b34abf49d |
| genomic_resource.yaml | 2.52 KB | a4ed45d574c114167941c6f9e85b47fc |
| genomic_resource_original.yaml | 2.42 KB | 58cbaad08efd6ea5909ed249af6e6bb6 |
| statistics/ |