TF_ChIP-seq_ENCSR000EWI
| Id: | TF_ChIP-seq/ENCSR000EWI |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000EWI [biosamplesummary="Homo sapiens K562" and target="SETDB1"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: output_type: IDR thresholded peaks audit_internal_action: Released analysis {ENCAN928BMR|/analyses/ENCAN928BMR/} has in progress subobject document {02cc9393-2f07-4677-a1dd-03e0c5b229b2|/documents/02cc9393-2f07-4677-a1dd-03e0c5b229b2/} audit_internal_action: Released analysis {ENCAN928BMR|/analyses/ENCAN928BMR/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_not_compliant: Processed alignments file {ENCFF495NJJ|/files/ENCFF495NJJ/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 9102690 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting SETDB1-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_not_compliant: Processed alignments file {ENCFF575NPK|/files/ENCFF575NPK/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 6403283 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting SETDB1-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) |
| Labels: |
|
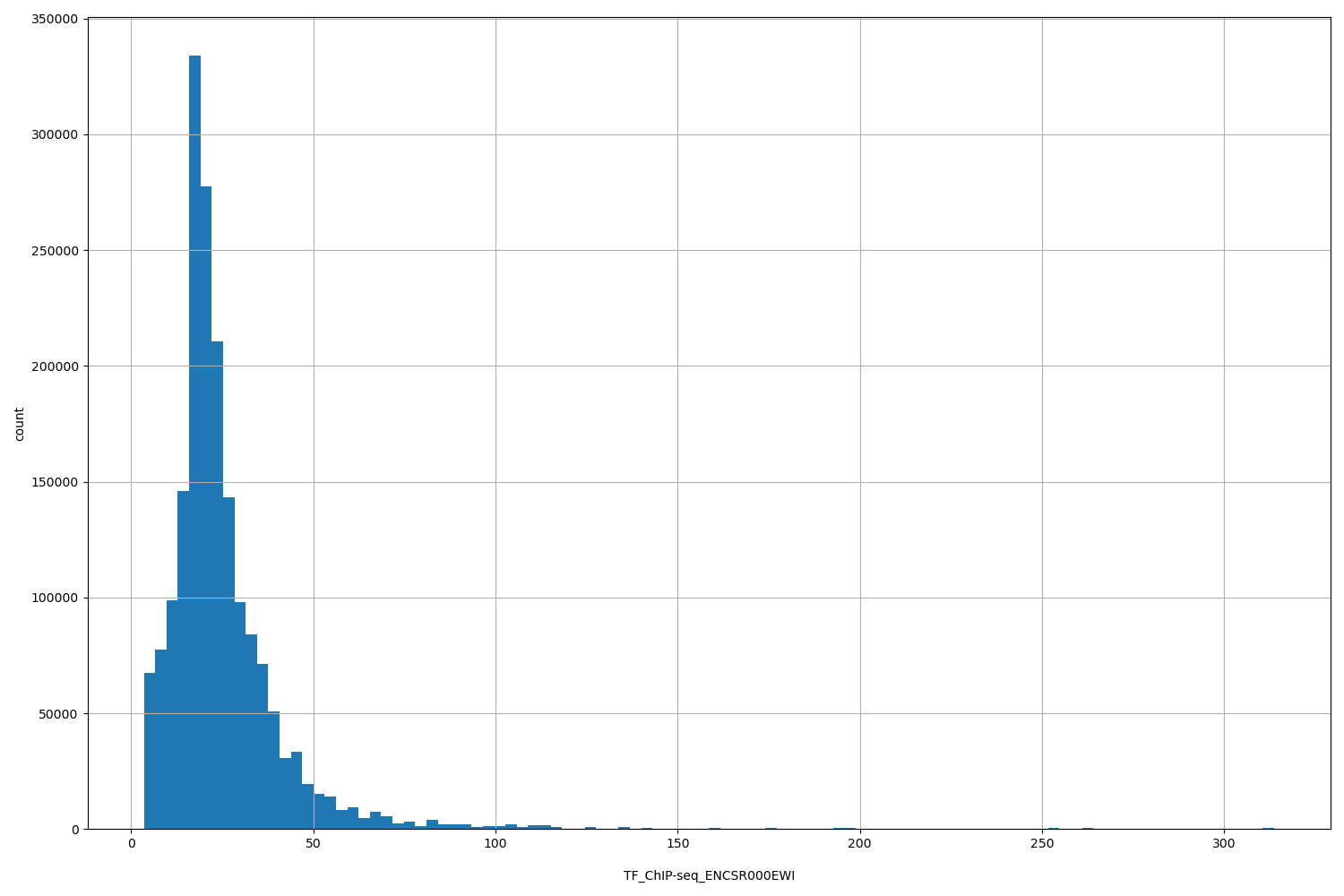
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000EWI | float |
TF_ChIP-seq_ENCSR000EWI |
TF_ChIP-seq ENCSR000EWI [biosample_summary="Homo sapiens K562" and target="SETDB1"]
|
 |
[3.55, 314] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF752EMN.bed.gz | 82.37 KB | bb903a17f51953a508b2c369028a8556 |
| ENCFF752EMN.bed.gz.dvc | 99.0 B | 1b92dff37cc651b50680840fbcdde282 |
| ENCFF752EMN.tabix.bed.gz | 62.26 KB | 5db95a8277e53ee3bb76c70e7ee1ce44 |
| ENCFF752EMN.tabix.bed.gz.dvc | 105.0 B | 3d7ecf1eb04d48575e03eae8a853ab70 |
| ENCFF752EMN.tabix.bed.gz.tbi | 41.59 KB | bb412814d6aeadbef61124f6ec60f6a1 |
| ENCFF752EMN.tabix.bed.gz.tbi.dvc | 109.0 B | ee11fbcd4efe46632a844ba74519e637 |
| genomic_resource.yaml | 2.58 KB | 0e76663e2f599a019ffbffcd14bdb752 |
| genomic_resource_original.yaml | 2.48 KB | 2ed39f6f92ec63ac5526bc6e79784c35 |
| statistics/ |