TF_ChIP-seq_ENCSR000EVR
TF_ChIP-seq ENCSR000EVR [biosample_summary="Homo sapiens HepG2" and target="ZNF274"]
| Id: | TF_ChIP-seq/ENCSR000EVR |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000EVR [biosamplesummary="Homo sapiens HepG2" and target="ZNF274"] |
| Description: |
status: released biological_replicates: summary: output_type: optimal IDR thresholded peaks audit_internal_action: derived_from is a list of files that were used to create a given file; for example, fastq file(s) will appear in the derived_from list of an alignments file. Processed file {ENCFF002CVA|/files/ENCFF002CVA/} is missing the requisite file specification in its derived_from list. |
| Labels: |
|
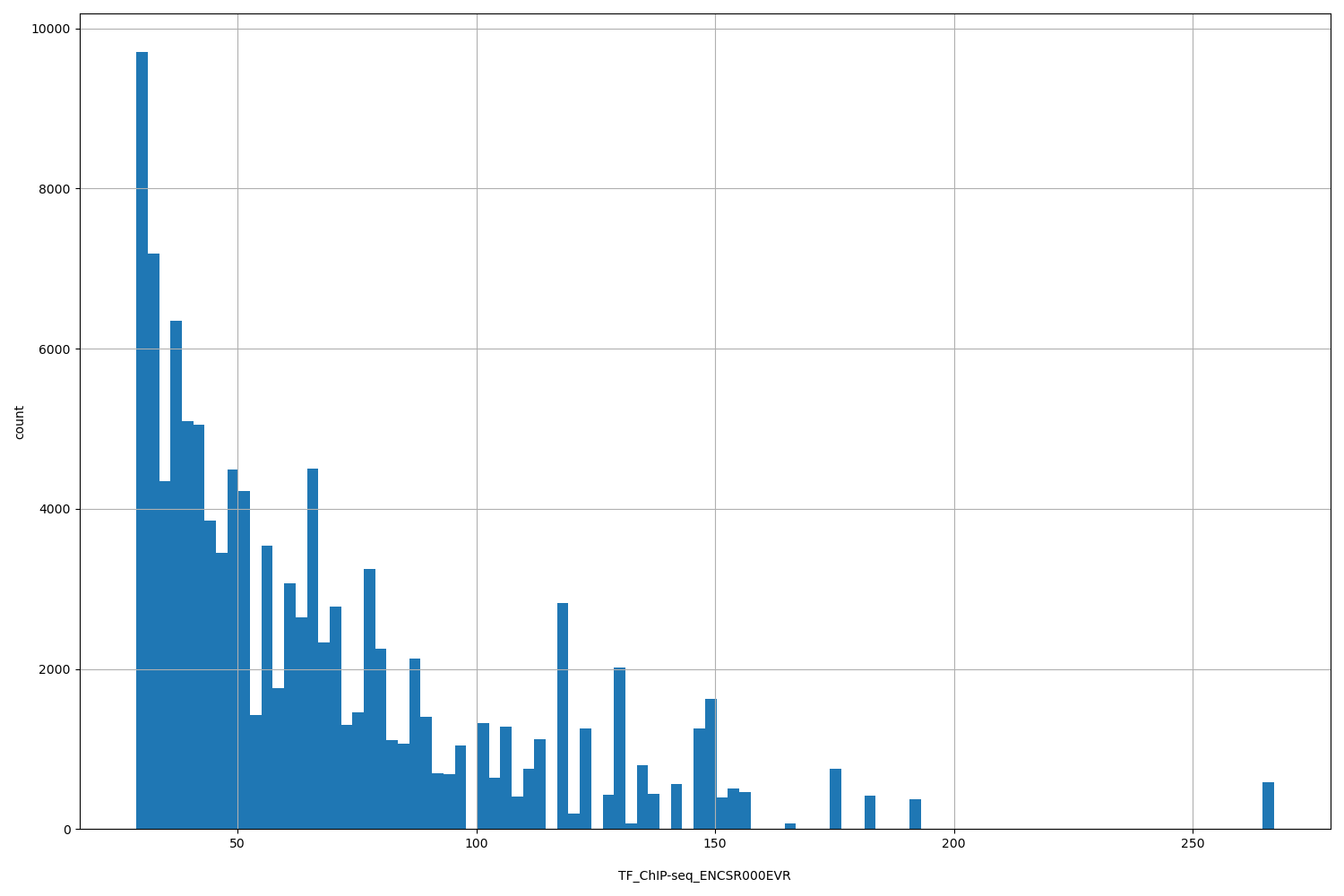
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000EVR | float |
TF_ChIP-seq_ENCSR000EVR |
TF_ChIP-seq ENCSR000EVR [biosample_summary="Homo sapiens HepG2" and target="ZNF274"]
|
 |
[28.8, 267] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF002CVA.bed.gz | 6.04 KB | 6d6e5c16bf06e488b8933b0b01da6aee |
| ENCFF002CVA.bed.gz.dvc | 98.0 B | 7b57036a6d8abba0c8044c95673bd4e0 |
| ENCFF002CVA.tabix.bed.gz | 4.21 KB | 75cbfb1d6cb6c3d2c7105ac7ba0871f2 |
| ENCFF002CVA.tabix.bed.gz.dvc | 104.0 B | f35a2b344fb8037231f7564abd3ccc11 |
| ENCFF002CVA.tabix.bed.gz.tbi | 4.56 KB | 7f702ee8c134b55bfde3e07a7c41a83d |
| ENCFF002CVA.tabix.bed.gz.tbi.dvc | 108.0 B | 46e5ecea37858a3dec718810025cbd4c |
| genomic_resource.yaml | 1.48 KB | 4397a884ecfe80bbc1f95fac52053345 |
| genomic_resource_original.yaml | 1.38 KB | 918264089e7d8d0d50b423005643fabb |
| statistics/ |