TF_ChIP-seq_ENCSR000EVQ
TF_ChIP-seq ENCSR000EVQ [biosample_summary="Homo sapiens HepG2" and target="TCF7L2"]
| Id: | TF_ChIP-seq/ENCSR000EVQ |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000EVQ [biosamplesummary="Homo sapiens HepG2" and target="TCF7L2"] |
| Description: |
status: released biological_replicates: summary: output_type: optimal IDR thresholded peaks audit_internal_action: derived_from is a list of files that were used to create a given file; for example, fastq file(s) will appear in the derived_from list of an alignments file. Processed file {ENCFF002CUX|/files/ENCFF002CUX/} is missing the requisite file specification in its derived_from list. |
| Labels: |
|
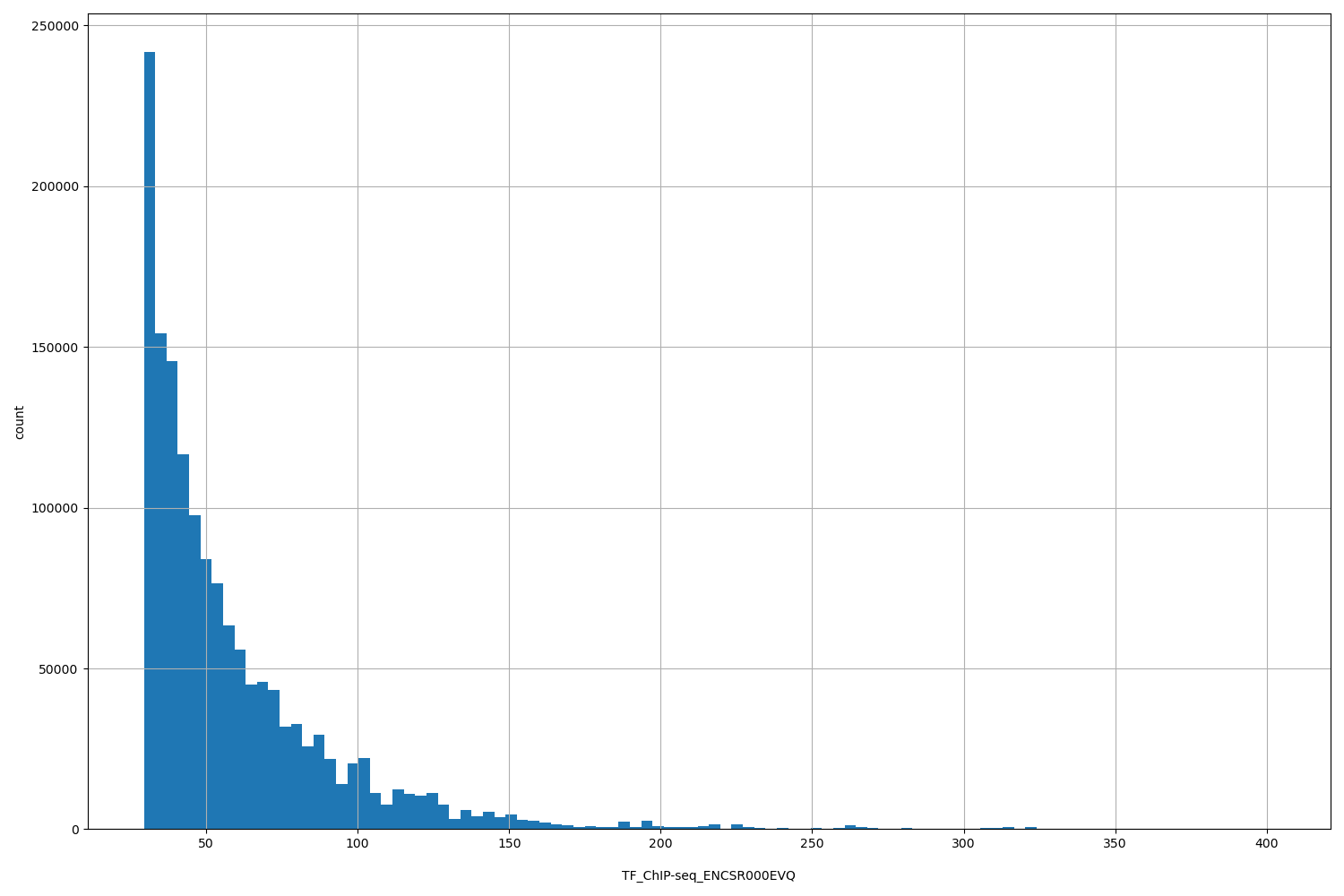
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000EVQ | float |
TF_ChIP-seq_ENCSR000EVQ |
TF_ChIP-seq ENCSR000EVQ [biosample_summary="Homo sapiens HepG2" and target="TCF7L2"]
|
 |
[29.4, 402] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF002CUX.bed.gz | 64.71 KB | 48217ad1de947d3aa04bab64adba7682 |
| ENCFF002CUX.bed.gz.dvc | 99.0 B | e42b75f28305538b703d77a375e57363 |
| ENCFF002CUX.tabix.bed.gz | 48.6 KB | a0d882509628d9bf3e6f15e9c3654f12 |
| ENCFF002CUX.tabix.bed.gz.dvc | 105.0 B | 587531ae2bf7bed08bded13b5e6029ee |
| ENCFF002CUX.tabix.bed.gz.tbi | 27.73 KB | 953a1584ee35fede0e881048f4ee74db |
| ENCFF002CUX.tabix.bed.gz.tbi.dvc | 109.0 B | 47641678990501a42683bef08b7d9b35 |
| genomic_resource.yaml | 1.48 KB | dcfe49a4199d0a00f06ee829ecae83f3 |
| genomic_resource_original.yaml | 1.38 KB | 381f77b2941a932c088771a8a6574745 |
| statistics/ |