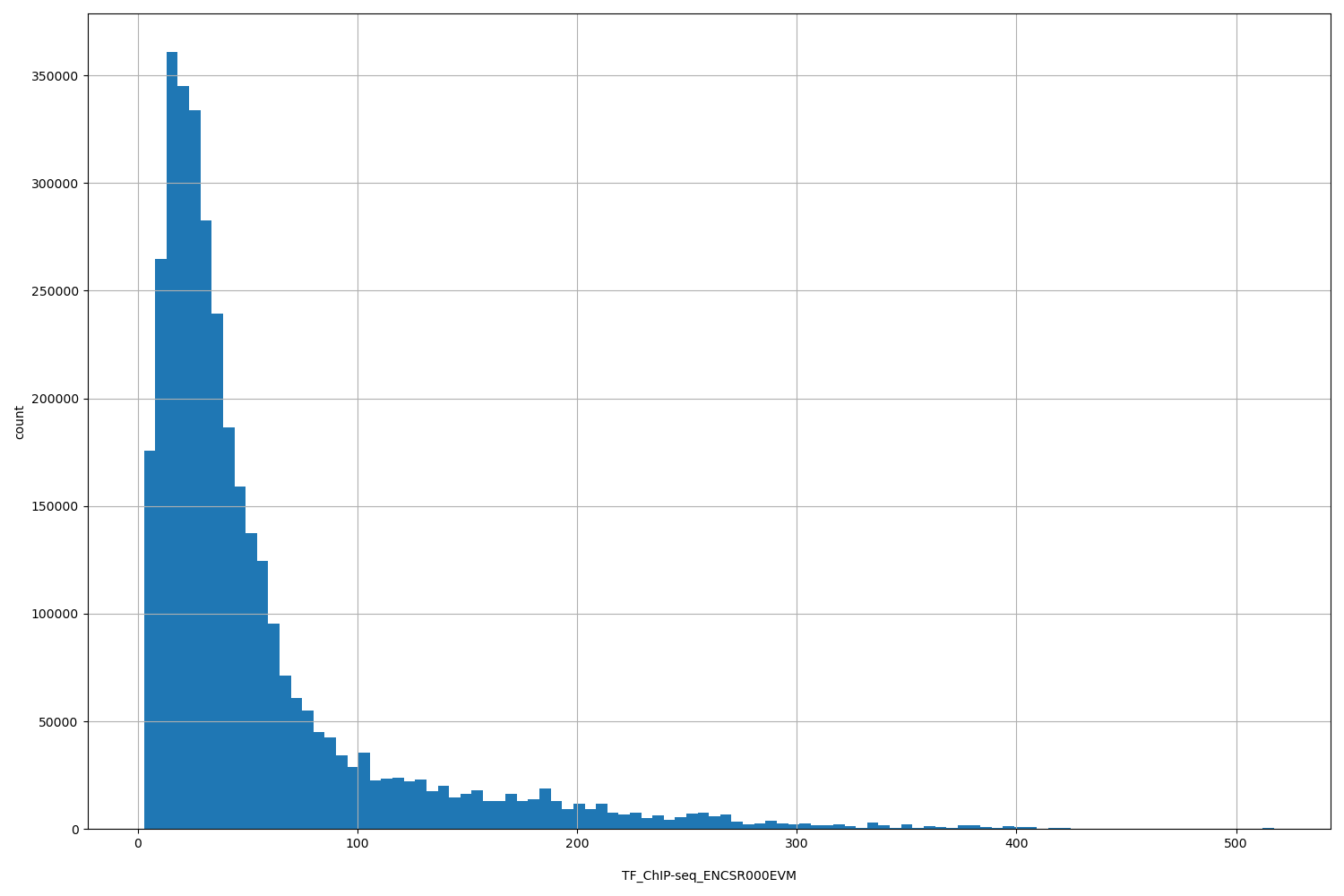
TF_ChIP-seq_ENCSR000EVM
| Id: | TF_ChIP-seq/ENCSR000EVM |
| Type: | position_score |
| Version: | 0 |
| Summary: |
TFChIP-seq ENCSR000EVM [biosamplesummary="Homo sapiens HeLa-S3 stably expressing E2F1" and target="E2F1"] |
| Description: |
status: released biological_replicates: Rep 1, Rep 2 summary: stably expressing N-terminal HA-tagged E2F1 output_type: IDR thresholded peaks audit_error: Processed alignments file {ENCFF250EHY|/files/ENCFF250EHY/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 3893368 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting E2F1-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_internal_action: Released analysis {ENCAN469TLP|/analyses/ENCAN469TLP/} has in progress subobject quality standard {encode4-tf-chip|/quality-standards/encode4-tf-chip/} audit_internal_action: Released analysis {ENCAN469TLP|/analyses/ENCAN469TLP/} has in progress subobject document {7b5c9829-16ab-4d13-b465-42fcd791d328|/documents/7b5c9829-16ab-4d13-b465-42fcd791d328/} audit_not_compliant: Processed alignments file {ENCFF900XOM|/files/ENCFF900XOM/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline has 9443505 usable fragments. The minimum ENCODE standard for each replicate in a ChIP-seq experiment targeting E2F1-human and investigated as a transcription factor is 10 million usable fragments. The recommended value is > 20 million, but > 10 million is acceptable. (See {ENCODE ChIP-seq data standards|/data-standards/chip-seq/} ) audit_warning: Replicate concordance in ChIP-seq experiments is measured by calculating IDR values (Irreproducible Discovery Rate). ENCODE processed IDR thresholded peaks files {ENCFF872RSG|/files/ENCFF872RSG/} processed by ChIP-seq ENCODE4 v1.6.1 GRCh38 pipeline have a rescue ratio of 1.02 and a self consistency ratio of 2.09. According to ENCODE standards, having both rescue ratio and self consistency ratio values < 2 is recommended, but having only one of the ratio values < 2 is acceptable. |
| Labels: |
|
| ID | Type | Default annotation | Description | Histogram | Range |
|---|---|---|---|---|---|
| TF_ChIP-seq_ENCSR000EVM | float |
TF_ChIP-seq_ENCSR000EVM |
TF_ChIP-seq ENCSR000EVM [biosample_summary="Homo sapiens HeLa-S3 stably expressing E2F1" and target="E2F1"]
|
 |
[2.79, 517] |
| Filename | Size | md5 |
|---|---|---|
| ENCFF872RSG.bed.gz | 186.26 KB | db396d86aa104c9f87b80f1584b9fbeb |
| ENCFF872RSG.bed.gz.dvc | 100.0 B | 859b9c70f87d3606c29aa2049c2d3b59 |
| ENCFF872RSG.tabix.bed.gz | 145.1 KB | cce069fbaaaac1b970b6bc5e91ae8e47 |
| ENCFF872RSG.tabix.bed.gz.dvc | 106.0 B | b7379eb3c27dd7fef33381a5937bb8aa |
| ENCFF872RSG.tabix.bed.gz.tbi | 76.7 KB | c6285e5b22a7a73658f64bdee7ab5cdb |
| ENCFF872RSG.tabix.bed.gz.tbi.dvc | 109.0 B | 88d25a35d0dce4eea6119bb9ce305c31 |
| genomic_resource.yaml | 3.18 KB | bfde2f4befb4a65eebdf18d603df79fc |
| genomic_resource_original.yaml | 3.07 KB | 0fe8b9ea1752743369f03de178cd68b2 |
| statistics/ |